Streaming Data on AWS: Amazon Kinesis Data Streams or Amazon MSK?
Posted by Gary A. Stafford in Analytics, AWS, Big Data, Serverless on April 23, 2023
Given similar functionality, what differences make one AWS-managed streaming service a better choice over the other?

Data streaming has emerged as a powerful tool in the last few years thanks to its ability to quickly and efficiently process large volumes of data, provide real-time insights, and scale and adapt to meet changing needs. As IoT, social media, and mobile devices continue to generate vast amounts of data, it has become imperative to have platforms that can handle the real-time ingestion, processing, and analysis of this data.
Key Differentiators
Amazon Kinesis Data Streams and Amazon Managed Streaming for Apache Kafka (Amazon MSK) are two managed streaming services offered by AWS. While both platforms offer similar features, choosing the right service largely depends on your specific use cases and business requirements.
Amazon Kinesis Data Streams
- Simplicity: Kinesis Data Streams is generally considered a less complicated service than Amazon MSK, which requires you to manage more of the underlying infrastructure. This can make setting up and managing your streaming data pipeline easier, especially if you have limited experience with Apache Kafka. Amazon MSK Serverless, which went GA in April 2022, is a cluster type for Amazon MSK that allows you to run Apache Kafka without managing and scaling cluster capacity. Unlike Amazon MSK provisioned, Amazon MSK Serverless greatly reduces the effort required to use Amazon MSK, making ‘Simplicity’ less of a Kinesis differentiator.
- Integration with AWS services: Kinesis Data Streams integrates well with other AWS services, such as AWS Lambda, Amazon S3, and Amazon OpenSearch. This can make building end-to-end data processing pipelines easier using these services.
- Low latency: Kinesis Data Streams is designed to deliver low-latency processing of streaming data, which can be important for applications that require near real-time processing.
- Predictable pricing: Kinesis Data Streams is generally considered to have a more predictable pricing model than Amazon MSK, based on instance sizes and hourly usage. With Kinesis Data Streams, you pay for the data you process, making estimating and managing fees easier (additional fees may apply).

Amazon MSK
- Compatibility with Apache Kafka: Amazon MSK may be a better choice if you have an existing Apache Kafka deployment or are already familiar with Kafka. Amazon MSK is a fully managed version of Apache Kafka, which you can use with existing Kafka applications and tools.
- Customization: With Amazon MSK, you have more control over the underlying cluster infrastructure, configuration, deployment, and version of Kafka, which means you can customize the cluster to meet your needs. This can be important if you have specialized requirements or want to optimize performance (e.g., high-volume financial trading, real-time gaming).
- Larger ecosystem: Apache Kafka has a large ecosystem of tools and integrations compared to Kinesis Data Streams. This can provide flexibility and choice when building and managing your streaming data pipeline. Some common tools include MirrorMaker, Kafka Connect, LinkedIn’s Cruise Control, kcat (fka kafkacat), Lenses, Confluent Schema Registry, and Appicurio Registry.
- Preference for Open Source: You may prefer the flexibility, transparency, pace of innovation, and interoperability of employing open source software (OSS) over proprietary software and services for your streaming solution.

Ultimately, the choice between Amazon Kinesis Data Streams and Amazon MSK will depend on your specific needs and priorities. Kinesis Data Streams might be better if you prioritize simplicity, integration with other AWS services, and low latency. If you have an existing Kafka deployment, require more customization, or need access to a larger ecosystem of tools and integrations, Amazon MSK might be a better fit. In my opinion, the newer Amazon MSK Serverless option lessens several traditional differentiators between the two services.
Scaling Capabilities
Amazon Kinesis Data Streams and Amazon MSK are designed to be scalable streaming services that can handle large volumes of data. However, there are some differences in their scaling capabilities.
Amazon Kinesis Data Streams
- Scalability: Kinesis Data Streams has two capacity modes, on-demand and provisioned. With the on-demand mode, Kinesis Data Streams automatically manages the shards to provide the necessary throughput based on the amount of data you process. This means the service can automatically adjust the number of shards based on the incoming data volume, allowing you to handle increased traffic without manually adjusting the infrastructure.
- Limitations: Per the documentation, there is no upper quota on the number of streams with the provisioned mode you can have in an account. A shard can ingest up to 1 MB of data per second (including partition keys) or 1,000 records per second for writes. The maximum size of the data payload of a record before base64-encoding is up to 1 MB. GetRecords can retrieve up to 10 MB of data per call from a single shard and up to 10,000 records per call. Each call to GetRecords is counted as one read transaction. Each shard can support up to five read transactions per second. Each read transaction can provide up to 10,000 records with an upper quota of 10 MB per transaction. Each shard can support a maximum total data read rate of 2 MB per second via GetRecords. If a call to GetRecords returns 10 MB, subsequent calls made within the next 5 seconds throw an exception.
- Cost: Kinesis Data Streams has two capacity modes — on-demand and provisioned — with different pricing models. With on-demand capacity mode, you pay per GB of data written and read from your data streams. You do not need to specify how much read and write throughput you expect your application to perform. With provisioned capacity mode, you select the number of shards necessary for your application based on its write and read request rate. There are additional fees
PUTPayload Units, enhanced fan-out, extended data retention, and retrieval of long-term retention data.
Amazon MSK
- Scalability: Amazon MSK is designed to be highly scalable and can handle millions of messages per second. With Amazon MSK provisioned, you can scale your Kafka cluster by adding or removing instances (brokers) and storage as needed. Amazon MSK can automatically rebalance partitions across instances. Alternately, Amazon MSK Serverless automatically provisions and scales capacity while managing the partitions in your topic, so you can stream data without thinking about right-sizing or scaling clusters.
- Flexibility: With Amazon MSK, you have more control over the underlying infrastructure, which means you can customize the deployment to meet your needs. This can be important if you have specialized requirements or want to optimize performance.
- Amazon MSK also offers multiple authentication methods. You can use IAM to authenticate clients and to allow or deny Apache Kafka actions. Alternatively, with Amazon MSK provisioned, you can use TLS or SASL/SCRAM to authenticate clients and Apache Kafka ACLs to allow or deny actions.
- Cost: Scaling up or down with Amazon MSK can impact the cost based on instance sizes and hourly usage. Therefore, adding more instances can increase the overall cost of the service. Pricing models for Amazon MSK and Amazon MSK Serverless vary.
Amazon Kinesis Data Streams and Amazon MSK are highly scalable services. Kinesis Data Streams can scale automatically based on the amount of data you process. At the same time, Amazon MSK allows you to scale your Kafka cluster by adding or removing instances and adding storage as needed. However, adding more shards with Kinesis can lead to a more manual process that can take some time to propagate and impact cost, while scaling up or down with Amazon MSK is based on instance sizes and hourly usage. Ultimately, the choice between the two will depend on your specific use case and requirements.
Throughput
Throughput can be measured in the maximum MB/s of data and the maximum number of records per second. The maximum throughput of both Amazon Kinesis Data Streams and Amazon MSK are not hard limits. Depending on the service, you can exceed these limits by adding more resources, including shards or brokers. Total maximum system throughput is affected by the maximum throughput of both upstream and downstream producing and consuming components.
Amazon Kinesis Data Streams
The maximum throughput of Kinesis Data Streams depends on the number of shards and the size of the data being processed. Each shard in a Kinesis stream can handle up to 1 MB/s of data input and up to 2 MB/s of data output, or up to 1,000 records per second for writes and up to 10,000 records per second for reads. When a consumer uses enhanced fan-out, it gets its own 2 MB/s allotment of read throughput, allowing multiple consumers to read data from the same stream in parallel without contending for read throughput with other consumers.
The maximum throughput of a Kinesis stream is determined by the number of shards you have multiplied by the maximum throughput per shard. For example, if you have a stream with 10 shards, the maximum throughput of the stream would be 10 MB/s for data input and 20 MB/s for data output, or up to 10,000 records per second for writes and up to 100,000 records per second for reads.
The maximum throughput is not a hard limit, and you can exceed these limits by adding more shards to your stream. However, adding more shards can impact the cost of the service, and you should consider the optimal shard count for your use case to ensure efficient and cost-effective processing of your data.
Amazon MSK
As discussed in the Amazon MSK best practices documentation, the maximum throughput of Amazon MSK depends on the number of brokers and the instance type of those brokers. Amazon MSK allows you to scale the number of instances in a Kafka cluster up or down based on your needs.
The maximum throughput of an Amazon MSK cluster depends on the number of brokers and the performance characteristics of the instance types you are using. Each broker in an Amazon MSK cluster can handle tens of thousands of messages per second, depending on the instance type and configuration. The actual throughput you can achieve will depend on your specific use case and the message size. The AWS blog post, Best practices for right-sizing your Apache Kafka clusters to optimize performance and cost, is an excellent reference.
The maximum throughput is not a hard limit, and you can exceed these limits by adding more brokers or upgrading to more powerful instances. However, adding more instances or upgrading to more powerful instances can impact the service’s cost. Therefore, consider your use case’s optimal instance count and type to ensure efficient and cost-effective data processing.
Writing Messages
Compatibility with multiple producers and consumers is essential when choosing a streaming technology. There are multiple ways to write messages to Amazon Kinesis Data Streams and Amazon MSK.
Amazon Kinesis Data Streams
- AWS SDK: Use the AWS SDK for your preferred programming language.
- Kinesis Producer Library (KPL): KPL is a high-performance library that allows you to write data to Kinesis Data Streams at a high rate. KPL handles all heavy lifting, including batching, retrying failed records, and load balancing across shards.
- Amazon Kinesis Data Firehose: Kinesis Data Firehose is a fully managed service that can ingest and transform streaming data in real-time. It can be used to write data to Kinesis Data Streams, as well as to other AWS services such as S3, Redshift, and Elasticsearch.
- Amazon Kinesis Data Analytics: Kinesis Data Analytics is a fully managed service that allows you to process and analyze streaming data in real-time. It can read data from Kinesis Data Streams, perform real-time analytics and transformations, and write the results to another Kinesis stream or an external data store.
- Kinesis Agent: Kinesis Agent is a standalone Java application that collects and sends data to Kinesis Data Streams. It can monitor log files or other data sources and automatically send data to Kinesis Data Streams as it is generated.
- Third-party libraries and tools: There are many third-party libraries and tools available for writing data to Kinesis Data Streams, including Apache Kafka Connect, Apache Storm, and Fluentd. These tools can integrate Kinesis Data Streams with existing data processing pipelines or build custom streaming applications.

Amazon MSK
- Kafka command line tools: The Kafka command line tools (e.g.,
kafka-console-producer.sh) can be used to write messages to a Kafka topic in an Amazon MSK cluster. These tools are part of the Kafka distribution and are pre-installed on the Amazon MSK broker nodes. - Kafka client libraries: You can use Kafka client libraries in your preferred programming language (e.g., Java, Python, C#) to write messages to an Amazon MSK cluster. These libraries provide a more flexible and customizable way to produce messages to Kafka topics.
- AWS SDKs: You can use AWS SDKs (e.g., AWS SDK for Java, AWS SDK for Python) to interact with Amazon MSK and write messages to Kafka topics. These SDKs provide a higher-level abstraction over the Kafka client libraries, making integrating Amazon MSK into your AWS infrastructure easier.
- Third-party libraries and tools: There are many third-party tools and frameworks, including Apache NiFi, Apache Camel, and Apache Beam. They provide Kafka connectors and producers, which can be used to write messages to Kafka topics in Amazon MSK. These tools can simplify the process of writing messages and provide additional features such as data transformation and routing.
Schema Registry
You can use AWS Glue Schema Registry with Amazon Kinesis Data Streams and Amazon MSK. AWS Glue Schema Registry is a fully managed service that provides a central schema repository for organizing, validating, and tracking the evolution of your data schemas. It enables you to store, manage, and discover schemas for your data in a single, centralized location.
With AWS Glue Schema Registry, you can define and register schemas for your data in the registry. You can then use these schemas to validate the data being ingested into your streaming applications, ensuring that the data conforms to the expected structure and format.
Both Kinesis Data Streams and Amazon MSK support the use of AWS Glue Schema Registry through the use of Apache Avro schemas. Avro is a compact, fast, binary data format that can improve the performance of your streaming applications. You can configure your streaming applications to use the registry to validate incoming data, ensuring that it conforms to the schema before processing.
Using AWS Glue Schema Registry can help ensure the consistency and quality of your data across your streaming applications and provide a centralized location for managing and tracking schema changes. Amazon MSK is also compatible with popular alternative schema registries, such as Confluent Schema Registry and RedHat’s open-source Apicurio Registry.

Stream Processing
According to TechTarget, Stream processing is a data management technique that involves ingesting a continuous data stream to quickly analyze, filter, transform, or enhance the data in real-time. Several leading stream processing tools are available, compatible with Amazon Kinesis Data Streams and Amazon MSK. Each tool with its own strengths and use cases. Some of the more popular tools include:
- Apache Flink: Apache Flink is a distributed stream processing framework that provides fast, scalable, and fault-tolerant data processing for real-time and batch data streams. It supports a variety of data sources and sinks and provides a powerful stream processing API and SQL interface. In addition, Amazon offers its managed version of Apache Flink, Amazon Kinesis Data Analytics (KDA), which is compatible with both Amazon Kinesis Data Streams and Amazon MSK.
- Apache Spark Structured Streaming: Apache Spark Structured Streaming is a stream processing framework that allows developers to build real-time stream processing applications using the familiar Spark API. It provides high-level APIs for processing data streams and supports integration with various data sources and sinks. Apache Spark is compatible with both Amazon Kinesis Data Streams and Amazon MSK. Spark Streaming is available as a managed service on AWS via AWS Glue Studio and Amazon EMR.
- Apache NiFi: Apache NiFi is an open-source data integration and processing tool that provides a web-based UI for building data pipelines. It supports batch and stream processing and offers a variety of processors for data ingestion, transformation, and delivery. Apache NiFi is compatible with both Amazon Kinesis Data Streams and Amazon MSK.
- Amazon Kinesis Data Firehose (KDA): Kinesis Data Firehose is a fully managed service that can ingest and transform streaming data in real time. It can be used to write data to Kinesis Data Streams, as well as to other AWS services such as S3, Redshift, and Elasticsearch. Kinesis Data Firehose is compatible with Amazon Kinesis Data Streams and Amazon MSK.
- Apache Kafka Streams (aka KStream): Apache Kafka Streams is a lightweight stream processing library that allows developers to build scalable and fault-tolerant real-time applications and microservices. KStreams integrates seamlessly with Amazon MSK and provides a high-level DSL for stream processing.
- ksqlDB: ksqlDB is a database for building stream processing applications on top of Apache Kafka. It is distributed, scalable, reliable, and real-time. ksqlDB combines the power of real-time stream processing with the approachable feel of a relational database through a familiar, lightweight SQL syntax. ksqlDB is compatible with Amazon MSK.
Several stream-processing tools are detailed in my recent two-part blog post, Exploring Popular Open-source Stream Processing Technologies.
Conclusion
Amazon Kinesis Data Streams and Amazon Managed Streaming for Apache Kafka (Amazon MSK) are managed streaming services. While they offer similar functionality, some differences might make one a better choice, depending on your use cases and experience. Ensure you understand your streaming requirements and each service’s capabilities before making a final architectural decision.
🔔 To keep up with future content, follow Gary Stafford on LinkedIn.
This blog represents my viewpoints and not those of my employer, Amazon Web Services (AWS). All product names, logos, and brands are the property of their respective owners.
Unlocking the Potential of Generative AI for Synthetic Data Generation
Posted by Gary A. Stafford in AI/ML, Cloud, Enterprise Software Development, Machine Learning, Python, Software Development, Technology Consulting on April 18, 2023
Explore the capabilities and applications of generative AI to create realistic synthetic data for software development, analytics, and machine learning

Introduction
Generative AI refers to a class of artificial intelligence algorithms capable of generating new data similar to a given dataset. These algorithms learn the underlying patterns and relationships in the data and use this knowledge to create new data consistent with the original dataset. Generative AI is a rapidly evolving field that has the potential to revolutionize the way we generate and use data.
Generative AI can generate synthetic data based on patterns and relationships learned from actual data. This ability to generate synthetic data has numerous applications, from creating realistic virtual environments for training and simulation to generating new data for machine learning models. In this article, we will explore the capabilities of generative AI and its potential to generate synthetic data, both directly and indirectly, for software development, data analytics, and machine learning.
Common Forms of Synthetic Data
According to AltexSoft, in their article Synthetic Data for Machine Learning: its Nature, Types, and Ways of Generation, common forms of synthetic data include:
- Tabular data: This type of synthetic data is often used to generate datasets that resemble real-world data in terms of structure and statistical properties.
- Time series data: This type of synthetic data generates datasets that resemble real-world time series data. It is commonly used when real-world time series data is unavailable or too expensive.
- Image and video data: This synthetic data is used to generate realistic images and videos for training machine learning models or simulations.
- Text data: This synthetic data generates realistic text for natural language processing tasks or for generating training data for machine learning models.
- Sound data: This synthetic data generates realistic sound for training machine learning models or simulations.
Synthetic Tabular Data Types
Synthetic tabular data refers to artificially generated datasets that resemble real-world tabular data in terms of structure and statistical properties. Tabular data is organized into rows and columns, like tables or spreadsheets. Some specific types of synthetic tabular data include:
- Financial data: Synthetic datasets that resemble real-world financial data such as bank transactions, stock prices, or credit card information.
- Customer data: Synthetic datasets that resemble real-world customer data, such as purchase history, demographic information, or customer behavior.
- Medical data: Synthetic datasets that resemble real-world medical data, such as patient records, medical test results, or treatment history.
- Sensor data: Synthetic datasets that resemble real-world sensor data such as temperature readings, humidity levels, or air quality measurements.
- Sales data: Synthetic datasets that resemble real-world sales data, such as sales transactions, product information, or customer behavior.
- Inventory data: Synthetic datasets that resemble real-world inventory data, such as stock levels, product information, or supplier information.
- Marketing data: Synthetic datasets that resemble real-world marketing data, such as campaign performance, customer behavior, or market trends.
- Human resources data: Synthetic datasets that resemble real-world human resources data, such as employee records, performance evaluations, or salary information.
Challenges with Creating Synthetic Data
According to sources including Towards Data Science, enov8, and J.P. Morgan, there are several challenges in creating synthetic data, including:
- Technical difficulty: Properly modeling complex real-world behaviors such as synthetic data is challenging, given available technologies.
- Biased behavior: The flexible nature of synthetic data makes it prone to potentially biased results.
- Privacy concerns: Care must be taken to ensure synthetic data does not reveal sensitive information.
- Quality of the data model: If the quality of the data model is not high, wrong conclusions can be reached.
- Time and effort: Synthetic data generation requires time and effort.
Difficult Patterns and Behaviors to Model
Many patterns and behaviors can be challenging to model in synthetic data, for example:
- Rare events: If certain events rarely occur in the real world, generating synthetic data that accurately reflects their distribution can be difficult.
- Complex relationships: Synthetic data generators may struggle to capture complex relationships between variables, such as non-linear interactions, feedback loops, or conditional dependencies.
- Contextual variability: Contextual factors, such as time, location, or individual differences, can have a significant impact on the distribution of data. Modeling this variability accurately can be challenging.
- Outliers and anomalies: Synthetic data generators may be unable to generate outliers or anomalies that are realistic and representative of the real world.
- Dynamic data: If the data is dynamic and changes over time, it can be challenging to generate synthetic data that captures these changes accurately.
- Unobserved variables: Sometimes, variables may be necessary for understanding the data distribution but are not directly observable. These variables can be challenging to model in synthetic data.
- Data bias: If the real-world data is biased in some way, such as over-representation of certain groups or under-representation of others, it can be challenging to generate synthetic data that is unbiased and representative of the population.
- Time-dependent patterns: If the data exhibits time-dependent patterns, such as seasonality or trends, it can be challenging to generate synthetic data that accurately reflects them.
- Spatial patterns: If the data has a spatial component, such as location data or images, it can be challenging to generate synthetic data that captures the spatial patterns realistically.
- Data sparsity: If the data is sparse or incomplete, it can be difficult to generate synthetic data that accurately reflects the distribution of the entire dataset.
- Human behavior: If the data involves human behavior, such as in social science or behavioral economics, it can be challenging to model the complex and nuanced behaviors of individuals and groups.
- Sensitive or confidential information: In some cases, the data may contain sensitive or confidential information that cannot be shared, making it challenging to generate synthetic data that preserves privacy while accurately reflecting the underlying distribution.
Overall, many patterns can be challenging to model accurately in synthetic data. It often requires careful consideration of the specific data characteristics and the synthetic data generation techniques’ limitations.
Easily Modeled Patterns and Behaviors
There are many simple and well-understood patterns that can be easily modeled in synthetic data, for example:
- Randomness: If the data is purely random, it can be easily generated using a random number generator or other simple techniques.
- Gaussian distribution: If the data follows a Gaussian or normal distribution, it can be generated using a Gaussian random number generator.
- Uniform distribution: If the data follows a uniform distribution, it can be easily generated using a uniform random number generator.
- Linear relationships: If the data follows a linear relationship between variables, it can be modeled using simple linear regression techniques.
- Categorical variables: If the data consists of categorical variables, such as gender or occupation, it can be generated using a categorical distribution.
- Text data: If the data consists of text, it can be generated using natural language processing techniques, such as language models or text generation algorithms.
- Time series: If the data consists of time series data, such as stock prices or weather data, it can be generated using time series models, such as Autoregressive integrated moving average (ARIMA) or Long short-term memory (LSTM).
- Seasonality: If the data exhibits seasonal patterns, such as higher sales data during holiday periods, it can be generated using seasonal time series models.
- Proportions and percentages: If the data consists of proportions or percentages, such as product sales distribution across different regions, it can be generated using beta or Dirichlet distributions.
- Multivariate normal distribution: If the data follows a multivariate normal distribution, it can be generated using a multivariate Gaussian random number generator.
- Networks: If the data consists of network or graph data, such as social networks or transportation networks, it can be generated using network models, such as Erdos-Renyi or Barabasi-Albert models.
- Binary data: If the data consists of binary data, such as whether a customer churned, it can be generated using a Bernoulli distribution.
- Geospatial data: If it involves geospatial data, such as the location of points of interest, it can be easily using geospatial models, such as point processes or spatial point patterns.
- Customer behaviors: If the data involves customer behaviors, such as browsing or purchase histories, it can be generated using customer journey models, such as Markov models.
In general, simple and well-understood patterns can be easily modeled using synthetic data techniques. In contrast, more complex and nuanced patterns may require more sophisticated modeling techniques and a deeper understanding of the underlying data characteristics.
Creating Synthetic Data with Generative AI
We can use many popular generative AI-powered tools to create synthetic data for testing applications, constructing analytics pipelines, and building machine learning models. Tools include OpenAI ChatGPT, Microsoft’s all-new Bing Chat, ChatSonic, Tabnine, GitHub Copilot, and Amazon CodeWhisperer. For more information on these tools, check out my recent blog post:
Accelerating Development with Generative AI-Powered Coding Tools
Explore six popular generative AI-powered tools, including ChatGPT, Copilot, CodeWhisperer, Tabnine, Bing, and…garystafford.medium.com
Let’s start with a simple example of generating synthetic sales data. Suppose we have created a new sales forecasting application for coffee shops that we need to test using synthetic data. We might start by prompting a generative AI tool like OpenAI’s ChatGPT for some data:
Create a CSV file with 25 random sales records for a coffee shop.
Each record should include the following fields:
- id (incrementing integer starting at 1)
- date (random date between 1/1/2022 and 12/31/2022)
- time (random time between 6:00am and 9:00pm in 1-minute increments)
- product_id (incrementing integer starting at 1)
- product
- calories
- price in USD
- type (drink or food)
- quantity (random integer between 1 and 3)
- amount (price * quantity)
- payment type (cash, credit, debit, or gift card)
The content and structure of a prompt can vary, and this can strongly influence ChatGPT’s response. Based on the above prompt, the results were accurate but not very useful for testing our application in this format. ChatGPT cannot create a physical CSV file. Furthermore, ChatGPT’s response length is limited; only about twenty records were returned. According to ChatGPT, in general, ChatGPT can generate responses of up to 2,048 tokens, the maximum output length allowed by the GPT-3 model.

Instead of outputting the actual synthetic data, we could ask ChatGPT to write a program that can, in turn, generate the synthetic data. This option is certainly more scalable. Let’s prompt ChatGPT to write a Python program to generate synthetic sales data with the same characteristics as before:
Create a Python3 program to generate 100 sales of common items
sold in a coffee shop. The data should be written to a CSV file
and include a header row. Each record should include the following fields:
- id (incrementing integer starting at 1)
- date (random date between 1/1/2022 and 12/31/2022)
- time (random time between 6:00am and 9:00pm in 1-minute increments)
- product_id (incrementing integer starting at 1)
- product
- calories
- price in USD
- type (drink or food)
- quantity (random integer between 1 and 3)
- amount (price * quantity)
- payment type (cash, credit, debit, or gift card)
Using a single concise prompt, ChatGPT generated a complete Python program, including code comments, to generate synthetic sales data. Unfortunately, given ChatGPT’s response size limitation, the coffee shop menu was limited to just six items. Reprompting for more items would result in the truncation of the output and, thus, the program, making it unrunnable. Instead, we could use an additional prompt to generate a longer Python list of menu items and combine the two pieces of code in our IDE. Regardless, we will still need to copy and paste the code into our IDE to review, debug, test, and run.

ChatGPT’s Python program, copied and pasted into VS Code, ran without modifications, and wrote 100 synthetic sales records to a CSV file!

Using IDE-based Generative AI Tools
Although generating synthetic data directly or snippets of code in chat-based generative AI tools are helpful for limited use cases, writing code in IDE gives us several advantages:
- Code does not need to be copied and pasted from external sources into an IDE
- Consecutive lines of code, method, and block code completion overcome the single response size limits of chat-based tools like OpenAI ChatGPT
- Code can be reused and adapted to evolving use cases over time
- Python interpreter and debugger or equivalent for other languages
- Automatic code formatting, linting, and code style enforcement
- Unit, integration, and functional testing
- Static code analysis (SCA)
- Vulnerability scanning
- IntelliSense for code completion
- Source code management (SCM) / version control
Let’s use the same techniques we used with ChatGPT, but from within an IDE to generate three types of synthetic data. We will choose Microsoft’s VS Code with GitHub Copilot and Python as our programming language.

Source Code
All the code examples shown in this post can be found on GitHub.
Example #1: Coffee Shop Sales Data
First, we will start by outlining the program’s objective using code comments on the top of our Python file. This detailed context helps us to clearly express our goal and enables Copilot to generate an accurate response.
# Write a program that creates synthetic sales data for a coffee shop.
# The program should accept a command line argument that specifies the number of records to generate.
# The program should write the sales data to a file called 'coffee_shop_sales_data.csv'.
# The program should contain the following functions:
# - main() function that calls the other functions
# - function that returns one random product from a list of dictionaries
# - function that returns a dictionary containing one sales record
# - function that writes the sales records to a file
Following the import statements also generated with the assistance of Copilot, we will write the first function to return a random product from a list of 25 products. Again, we will use code comments as a prompt to generate the code. Copilot was able to generate 100% of the function’s code from the comments.
# Write a function to create list of dictionaries.
# The list of dictionaries should contain 15 drink items and 10 food items sold in a coffee shop.
# Include the product id, product name, calories, price, and type (Food or Drink).
# Capilize the first letter of each product name.
# Return a random item from the list of dictionaries.
Below is an example of Copilot’s ability to generate complete lines of code. Ultimately, it generated 100% of the function including choosing the items sold in a coffee shop, with a reasonable price and caloric count. Copilot is not limited to just understanding code.

Next, we will write a function to return a random sales record.
# Write a function to return a random sales record.
# The record should be a dictionary with the following fields:
# - id (an incrementing integer starting at 1)
# - date (a random date between 1/1/2022 and 12/31/2022)
# - time (a random time between 6:00am and 9:00pm in 1 minute increments)
# - product_id, product, calories, price, and type (from the get_product function)
# - quantity (a random integer between 1 and 3)
# - amount (price * quantity)
# - payment type (Cash, Credit, Debit, Gift Card, Apple Pay, Google Pay, or Venmo)
Again, Copilot generated 100% of the function’s code as a single block based on the code comments.

Lastly, we will create a function to write the sales data into a CSV file using Copilot’s help.
# Write a function to write the sales records to a CSV file called 'coffee_shop_sales.csv'.
# Use an input parameter to specify the number of records to write.
# The CSV file must have a header row and be comma delimited.
# All string values must be enclosed in double quotes.
Again below, we see an example of Copilot’s ability to generate an entire Python function. I needed to correct a few problems with the generated code. First, there was a lack of quotes for string values, which I added to the function (quotechar='"', quoting=csv.QUOTE_NONNUMERIC). Also, the function was missing a key line of code, sale = get_sales_record(), which would have caused the code to fail. Remember, just because the code was generated does not mean it is correct.

Here is the complete program that creates synthetic sales data for a coffee shop with Copilot assistance:
Copilot generated an astounding 80–85% of the program’s final code. The initial program took 10–15 minutes to write using code comments. I then added a few new features, including the ability to pass in the record count on the command line and the hash-based transaction id, which took another 5 minutes. Finally, I used GitHub Code Brushes to optimize the code and generate the Python docstrings, and Black Formatter and Flake8 extensions to format and lint, all of which took less than 5 minutes. With testing and debugging, the total time was about 25–30 minutes.
The most significant difference with Copilot was that I never had to leave the IDE to look up code references or find existing sales datasets or even a coffee shop menu to duplicate. The code, as well as the list of products, price, calories, and product type, were all generated by Copilot.
To make this example more realistic, you could use Copilot’s assistance to write algorithms capable of reflecting daily, weekly, and seasonal variations in product choice and sales volumes. This might include simulating increased sales during the busy morning rush hour or a preference for iced drinks in the summer months versus hot drinks during the winter months.
Here is an example of the synthetic sales data output by the example application:
Example #2: Residential Address Data
We could use these same techniques to generate a list of residential addresses. To start, we can prompt Copilot for the values in a list of common street names and street types in the United States:
# Write a function that creates a list of common street names
# in the United States, in alphabetical order.
# List should be in alphabetical order. Each name should be unique.
# Return a random street name.
def get_street_name():
street_names = [
"Ash", "Bend", "Bluff", "Branch", "Bridge", "Broadway", "Brook", "Burg",
"Bury", "Canyon", "Cape", "Cedar", "Cove", "Creek", "Crest", "Crossing",
"Dale", "Dam", "Divide", "Downs", "Elm", "Estates", "Falls", "Fifth",
"First", "Fork", "Fourth", "Glen", "Green", "Grove", "Harbor", "Heights",
"Hickory", "Hill", "Hollow", "Island", "Isle", "Knoll", "Lake", "Landing",
...
]
return random.choice(street_names)
# Write a function that creates a list of common street types
# in the United States, in alphabetical order.
# List should be in alphabetical order. Each name should be unique.
# Return a random street type.
def get_street_type():
street_types = [
"Alley", "Avenue", "Bend", "Bluff", "Boulevard", "Branch", "Bridge", "Brook",
"Burg", "Circle", "Commons", "Court", "Drive", "Highway", "Lane", "Parkway",
"Place", "Road", "Square", "Street", "Terrace", "Trail", "Way"
]
return random.choice(street_types)
Next, we can create a function that returns a property type based on a categorical distribution of common residential property types with the prompt:
# Write a function to return a random property type.
# Accept a random value between 0 and 1 as an input parameter.
# The function must return one of the following values based on the %:
# 63% Single-family, 26% Multi-family, 4% Condo,
# 3% Townhouse, 2% Mobile home, 1% Farm, 1% Other.
Again, Copilot generated 100% of the function’s code as a single block based on the code comments.

Additionally, we could have Copilot help us generate a list of the 50 largest cities in the United States with state, zip code, and population, with the prompt:
# Write a function to returns the 50 largest cities in the United States.
# List should be sorted in descending order by population.
# Include the city, state abbreviation, zip code, and population.
# Return a list of dictionaries.
Once again, Copilot generated 100% of the function’s code using a combination of single lines and code blocks based on the code comments.

When randomly choosing a city, we can use a categorical distribution of populations of all the cities to control the distribution of cities in the final synthetic dataset. For example, there will be more addresses in larger cities like New York City or Los Angeles than in smaller cities like Buffalo or Virginia Beach, with the prompt:
# Write a function that calculates the total population of the list of cities.
# Add a 'pcnt_of_total_population' and 'pcnt_running_total' columns to list.
# Returns a sorted list of cities by population.
Here is the complete program that creates synthetic US-based address data with Copilot assistance:
To make this example more realistic, you could use Copilot’s assistance to write algorithms capable of accurately reflecting assessed property values based on the type of residence and the zip code.
Here is an example of the synthetic US-based residential address data output by the example application:
Example #3: Demographic Data
We could use the same techniques again to generate synthetic demographic data. With the assistance of Copilot, we can write functions that randomly return typical feminine or masculine first names (forenames) and common last names (surnames) found in the United States, for this use case.
# Write a function that generates a list of common feminine first names in the United States.
# List should be in alphabetical order.
# Each name should be unique.
# Return random first name.
def get_first_name_feminine():
first_name_feminine = [
"Alice", "Amanda", "Amy", "Angela", "Ann", "Anna", "Barbara", "Betty",
"Brenda", "Carol", "Carolyn", "Catherine", "Christine", "Cynthia", "Deborah", "Debra",
"Diane", "Donna", "Doris", "Dorothy", "Elizabeth", "Frances", "Gloria", "Heather",
"Helen", "Janet", "Jennifer", "Jessica", "Joyce", "Julie", "Karen", "Kathleen", "Kimberly",
]
return random.choice(first_name_feminine)
With the assistance of Copilot, we can also write functions that return demographic information, such as age, gender, race, marital status, religion, and political affiliation. Similar to the previous sales data example, we can influence the final synthetic dataset based on categorical distributions of different demographic categories, for instance, with the prompt:
# Write a function that returns a person's martial status.
# Accept a random value between 0 and 1 as an input parameter.
# The function must return one of the following values based on the %:
# 50% Married, 33% Single, 17% Unknown.
def get_martial_status(rnd_value):
if rnd_value < 0.50:
return "Married"
elif rnd_value < 0.83:
return "Single"
else:
return "Unknown"
By altering the categorical distributions, we can quickly alter the resulting synthetic dataset to reflect differing demographic characteristics: an older or younger population, the predominance of a single race, religious affiliation, or marital status, or the ratio of males to females.
Next, we can use a Gaussian distribution (aka normal distribution) to return the year of birth in a bell-shaped curve, given a mean year and a standard deviation, using Python’s random.normalvariate function.
# Write a function that generates a normal distribution of date of births.
# with a mean year of 1975 and a standard deviation of 10.
# Return random date of birth as a string in the format YYYY-MM-DD
def get_dob():
day_of_year = random.randint(1, 365)
year_of_birth = int(random.normalvariate(1975, 10))
dob = date(int(year_of_birth), 1, 1) + timedelta(day_of_year - 1)
dob = dob.strftime("%Y-%m-%d")
return dob
Here is the complete program that creates synthetic demographic data with Copilot’s assistance:
To make this example more realistic, you could use Copilot’s assistance to write algorithms capable of more accurately representing the nuanced associations and correlations between age, gender, race, marital status, religion, and political affiliation.
Here is an example of the synthetic demographic data output by the example application:
Generative AI Tools for Unit Testing
In addition to writing code and documentation, a common use of generative AI code assistants like Copilot is unit tests. For example, we can create unit tests for each function in our coffee shop sales data code generator, using the same method of prompting with code comments.

Conclusion
In this post, we learned how Generative AI could assist us in creating synthetic data for software development, analytics, and machine learning. The examples herein generated data using simple techniques. Using advanced modeling techniques, we could generate increasingly complex, realistic synthetic data.
To learn about other ways Generative AI can be used to assist in writing code, please read my previous article, Ten Ways to Leverage Generative AI for Development on AWS.
🔔 To keep up with future content, follow Gary Stafford on LinkedIn.
This blog represents my viewpoints and not those of my employer, Amazon Web Services (AWS). All product names, logos, and brands are the property of their respective owners.
Navigating the World of Generative AI: A Guide to Essential Terminology
Posted by Gary A. Stafford in AI/ML, Machine Learning, Technology Consulting on April 9, 2023
Learn the essential terms and concepts you need to know to navigate the rapidly evolving world of generative AI

Generative AI is a fascinating and rapidly evolving field that has the potential to transform the way we interact with technology. However, with so much buzz and hype surrounding this topic, making sense of it all can be challenging. In this article, we’ll cut through the noise and gain a clear understanding of the essential terminology you need to know to navigate the world of generative AI.
According to a variety of sources, including McKinsey & Company and Vox Media, the critical difference between generative AI and other emerging technologies is that millions of people can — and already are — using it to create new content, such as text, photos, video, code, and 3D renderings, from data it is trained on. Recent breakthroughs in the field have the potential to drastically change the way we approach content creation. This has led to widespread excitement and some understandable apprehension about the potential for generative AI to impact virtually every aspect of society and disrupt industries, including media and entertainment, healthcare and life sciences, education, advertising, legal services, and finance.
Even if your current role is not in technology, it is highly likely that generative AI will have a direct impact on both your personal and professional life.
Gary Stafford
Even if your current role is not in technology, it is highly likely that generative AI will have a direct impact on both your personal and professional life. Familiarizing yourself with basic terminology related to generative AI can help you better comprehend the discussions on social media and in the news.
Terminology
Let’s explore the following terminology (in alphabetical order):
- Artificial General Intelligence (AGI)
- Artificial Intelligence (AI)
- ChatGPT
- DALL·E
- Deep Learning (DL)
- Generative AI
- Generative Pre-trained Transformer (GPT)
- Intelligence Amplification (IA)
- Large Language Model (LLM)
- Machine Learning (ML)
- Neural Network
- OpenAI
- Prompt Engineering
- Reinforcement Learning with Human Feedback (RLHF).
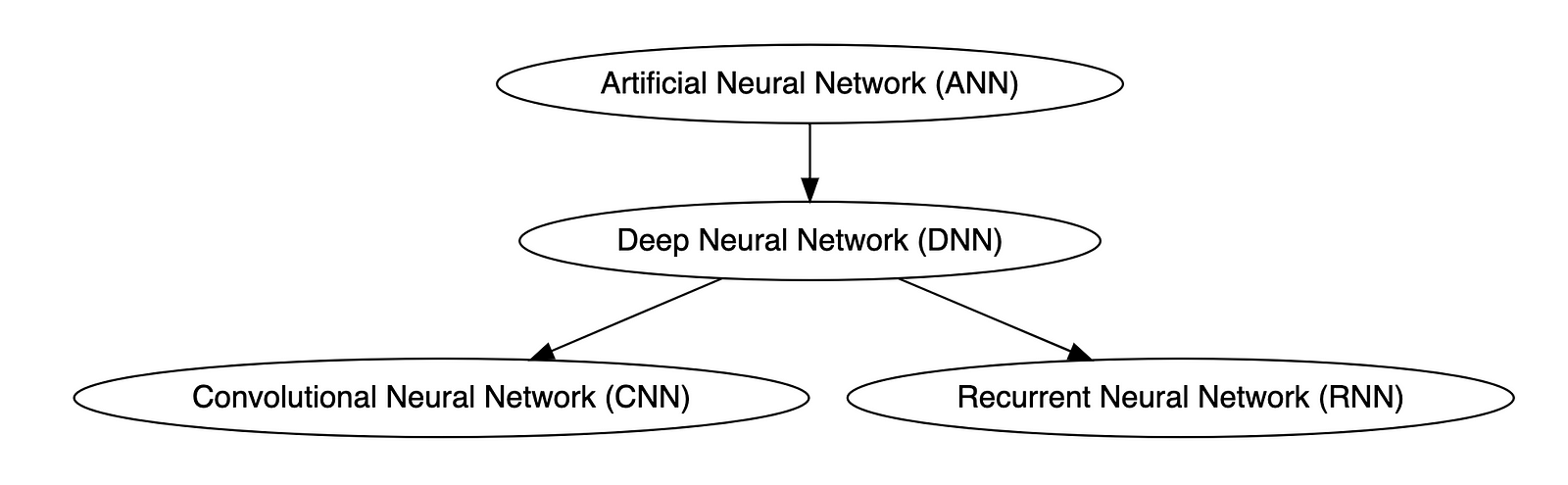
Below is a knowledge graph, created with OpenAI ChatGPT, showing the approximate relationships between the post’s terms.

Artificial General Intelligence (AGI)
According to the all-new Bing Chat, based on ChatGPT, artificial general intelligence (AGI) is the ability of an intelligent agent to understand or learn any intellectual task that human beings or other animals can. It is a primary goal of some artificial intelligence research and a common topic in science fiction and Futurism. According to Forbes, which also prompted ChatGPT, Artificial General Intelligence (AGI) refers to a theoretical type of artificial intelligence that possesses human-like cognitive abilities, such as the ability to learn, reason, solve problems, and communicate in natural language.
Eliezer Yudkowsky is an American researcher, writer, and philosopher on the topic of AI. The podcast Eliezer Yudkowsky: Dangers of AI and the End of Human Civilization, by prominent MIT Research Scientist Lex Fridman, explores various aspects of artificial general intelligence against the backdrop of the recent release of OpenAI’s GPT-4.
Artificial Intelligence (AI)
According to the Brookings Institute, AI is generally thought to refer to machines that respond to stimulation consistent with traditional responses from humans, given the human capacity for contemplation, judgment, and intention. Similarly, according to the U.S. Department of State, the term artificial intelligence refers to a machine-based system that can, for a given set of human-defined objectives, make predictions, recommendations, or decisions influencing real or virtual environments.

ChatGPT
ChatGPT, according to ChatGPT, is a large language model developed by OpenAI. I was trained on a massive dataset of human-written text using a deep neural network (DNN) architecture called GPT (Generative Pre-trained Transformer). Its purpose is to generate human-like responses to questions and prompts, engage in conversations, and perform various language-related tasks. It is a virtual assistant capable of understanding and generating natural language.
DALL·E
According to Wikipedia, DALL·E is a deep learning model developed by OpenAI to generate digital images from natural language descriptions, called prompts. DALL·E is a portmanteau of the names of the animated robot Pixar character WALL-E and the Spanish surrealist artist Salvador Dalí. According to OpenAI, DALL·E is an AI system that can create realistic images and art from a description in natural language. OpenAI introduced DALL·E in January 2021. One year later, in April 2022, they announced their newest system, DALL·E 2, which generates more realistic and accurate images with 4x greater resolution. DALL·E 2 can create original, realistic images and art from a text description. It can combine concepts, attributes, and styles.
Deep Learning (DL)
According to IBM, deep learning is a subset of machine learning (ML), which is essentially a neural network with three or more layers. These neural networks attempt to simulate the behavior of the human brain — albeit far from matching its ability — allowing it to “learn” from large amounts of data. While a neural network with a single layer can still make approximate predictions, additional hidden layers can help to optimize and refine for accuracy.
Generative AI
According to Wikipedia, generative artificial intelligence (AI), aka generative AI, is a type of AI system capable of generating text, images, or other media in response to prompts. Generative AI systems use generative models such as large language models (LLMs) to statistically sample new data based on the training data set used to create them.
Generative Pre-trained Transformer (GPT)
According to ChatGPT, Generative Pre-trained Transformer (GPT) is a deep learning architecture used for natural language processing (NLP) tasks, such as text generation, summarization, and question-answering. It uses a transformer neural network architecture with a self-attention mechanism, allowing the model to understand each word’s context in a sentence or text. The success of GPT models lies in their ability to generate natural-sounding and coherent text similar to human-written language. The term “pre-trained” refers to the fact that the model is trained on large amounts of unlabeled text data, such as books or web pages, to learn general language patterns and features, before being fine-tuned on smaller labeled datasets for specific tasks.
According to ZDNET, GPT-4, announced on March 14, 2023, is the newest version of OpenAI’s language model systems. Its previous version, GPT 3.5, powered the company’s wildly popular ChatGPT chatbot when it launched in November 2022. According to OpenAI, GPT-4 is the latest milestone in OpenAI’s effort to scale up deep learning. GPT-4 is a large multimodal model (accepting image and text inputs, emitting text outputs) that, while less capable than humans in many real-world scenarios, exhibits human-level performance on various professional and academic benchmarks.
Intelligence Amplification
According to Wikipedia, intelligence amplification (IA) (aka cognitive augmentation, machine-augmented intelligence, or enhanced intelligence) refers to the effective use of information technology in augmenting human intelligence. Similarly, Harvard Business Review describes intelligence amplification as the use of technology to augment human intelligence. And a paradigm shift is on the horizon, where new devices will offer less intrusive, more intuitive ways to amplify our intelligence.
In his latest book, Impromptu: Amplifying Our Humanity Through AI, co-authored by ChatGPT-4, Greylock general partner Reid Hoffman discusses the subject of intelligence amplification and AI’s ability to amplify human ability. The topic was also explored in Hoffman’s interview with OpenAI CEO Sam Altman on Greylock’s podcast series AI Field Notes.
Large Language Model (LLM)
According to Wikipedia, a large language model (LLM) is a language model consisting of a neural network with many parameters (typically billions of weights or more), trained on large quantities of unlabelled text using self-supervised learning. Though the term large language model has no formal definition, it often refers to deep learning models having a parameter count on the order of billions or more.
Machine Learning (ML)
According to MIT, machine learning (ML) is a subfield of artificial intelligence, which is broadly defined as the capability of a machine to imitate intelligent human behavior. Artificial intelligence systems are used to perform complex tasks in a way that is similar to how humans solve problems. The function of a machine learning system can be descriptive, meaning that the system uses the data to explain what happened; predictive, meaning the system uses the data to predict what will happen; or prescriptive, meaning the system will use the data to make suggestions about what action to take.
Neural Network
According to MathWorks, a neural network (aka artificial neural network or ANN) is an adaptive system that learns by using interconnected nodes or neurons in a layered structure that resembles a human brain. A neural network can learn from data to be trained to recognize patterns, classify data, and forecast future events. Similarly, according to AWS, a neural network is a method in artificial intelligence that teaches computers to process data in a way that is inspired by the human brain. It is a type of machine learning process called deep learning that uses interconnected nodes or neurons in a layered structure that resembles the human brain. It creates an adaptive system that computers use to learn from their mistakes and improve continuously.
Types of Neural Networks
Deep neural networks (DNNs) are improved versions of conventional artificial neural networks (ANNs) with multiple layers. While ANNs consist of one or two hidden layers to process data, DNNs contain multiple layers between the input and output layers. Convolutional neural networks (CNNs) are another kind of DNN. CNNs have a convolution layer, which uses filters to convolve an area of input data into a smaller area, detecting important or specific parts within the area. Recurrent neural networks (RNNs) can be considered a type of DNN. DNNs are neural networks with multiple layers between the input and output layers. RNNs can have multiple layers and can be used to process sequential data, making them a type of DNN.
OpenAI
OpenAI is a San Francisco-based AI research and deployment company whose mission is to “ensure that artificial general intelligence benefits all of humanity.” According to Wikipedia, OpenAI was founded in 2015 by current CEO Sam Altman, Greylock general partner Reid Hoffman, Y Combinator founding partner Jessica Livingston, Elon Musk, Ilya Sutskever, Peter Thiel, and others. OpenAI’s current products include GPT-4, DALL·E 2, Whisper, ChatGPT, and OpenAI Codex.

Prompt Engineering
According to Cohere, prompting (aka prompt engineering) is at the heart of working with LLMs. The prompt provides context for the text we want the model to generate. The prompts we create can be anything from simple instructions to more complex pieces of text, and they are used to encourage the model to produce a specific type of output. Cohere’s Generative AI with Cohere blog post series is an excellent resource on the topic of Prompting. Similarly, according to Dataconomy, using prompts to get the desired result from an AI tool is known as AI prompt engineering. A prompt can be a statement or a block of code, but it can also just be a string of words. Similar to how you may prompt a person as a starting point for writing an essay, you can use prompts to teach an AI model to produce the desired results when given a specific task.
Reinforcement Learning with Human Feedback (RLHF)
According to Scale AI in their blog, Why is ChatGPT so good?, instead of simply predicting the next word(s), large language models (LLMs) can now follow human instructions and provide useful responses. These advancements are made possible by fine-tuning them with specialized instruction datasets and a technique called reinforcement learning with human feedback (RLHF). Similarly, according to Hugging Face, RLHF (aka RL from human preferences) uses methods from reinforcement learning to directly optimize LLMs with human feedback. RLHF has enabled language models to begin to align a model trained on a geprogneral corpus of text data to that of complex human values.
Ready for More?
Mastered all the terminology, ready for more? Here are some additional generative AI terms for you to learn:
- Alignment (AI Alignment, Aligned AI)
- Attention (Self-attention)
- Backpropagation
- Embeddings
- Foundational Model
- Next-Token Predictors
- Parameter
- Reinforcement Learning (RL)
- Self-Supervised Learning (SSL)
- Temperature
- Tokenization
- Transformer
- Vector
- Weight
🔔 To keep up with future content, follow Gary Stafford on LinkedIn.
This blog represents my viewpoints and not those of my employer, Amazon Web Services (AWS). All product names, logos, and brands are the property of their respective owners.
Ten Ways to Leverage Generative AI for Development on AWS
Posted by Gary A. Stafford in AI/ML, AWS, Bash Scripting, Big Data, Build Automation, Client-Side Development, Cloud, DevOps, Enterprise Software Development, Kubernetes, Python, Serverless, Software Development, SQL on April 3, 2023
Explore ten ways you can use Generative AI coding tools to accelerate development and increase your productivity on AWS

Generative AI coding tools are a new class of software development tools that leverage machine learning algorithms to assist developers in writing code. These tools use AI models trained on vast amounts of code to offer suggestions for completing code snippets, writing functions, and even entire blocks of code.
Quote generated by OpenAI ChatGPT
Introduction
Combining the latest Generative AI coding tools with a feature-rich and extensible IDE and your coding skills will accelerate development and increase your productivity. In this post, we will look at ten examples of how you can use Generative AI coding tools on AWS:
- Application Development: Code, unit tests, and documentation
- Infrastructure as Code (IaC): AWS CloudFormation, AWS CDK, Terraform, and Ansible
- AWS Lambda: Serverless, event-driven functions
- IAM Policies: AWS IAM policies and Amazon S3 bucket policies
- Structured Query Language (SQL): Amazon RDS, Amazon Redshift, Amazon Athena, and Amazon EMR
- Big Data: Apache Spark and Flink on Amazon EMR, AWS Glue, and Kinesis Data Analytics
- Configuration and Properties files: Amazon MSK, Amazon EMR, and Amazon OpenSearch
- Apache Airflow DAGs: Amazon MWAA
- Containerization: Kubernetes resources, Helm Charts, Dockerfiles for Amazon EKS
- Utility Scripts: PowerShell, Bash, Shell, and Python
Choosing a Generative AI Coding Tool
In my recent post, Accelerating Development with Generative AI-Powered Coding Tools, I reviewed six popular tools: ChatGPT, Copilot, CodeWhisperer, Tabnine, Bing, and ChatSonic.
For this post, we will use GitHub Copilot, powered by OpenAI Codex, a new AI system created by OpenAI. Copilot suggests code and entire functions in real-time, right from your IDE. Copilot is trained in all languages that appear in GitHub’s public repositories. GitHub points out that the quality of suggestions you receive may depend on the volume and diversity of training data for that language. Similar tools in this category are limited in the number of languages they support compared to Copilot.

Copilot is currently available as an extension for Visual Studio Code, Visual Studio, Neovim, and JetBrains suite of IDEs. The GitHub Copilot extension for Visual Studio Code (VS Code) already has 4.8 million downloads, and the GitHub Copilot Nightly extension, used for this post, has almost 280,000 downloads. I am also using the GitHub Copilot Labs extension in this post.

Ten Ways to Leverage Generative AI
Take a look at ten examples of how you can use Generative AI coding tools to increase your development productivity on AWS. All the code samples in this post can be found on GitHub.
1. Application Development
According to GitHub, trained on billions of lines of code, GitHub Copilot turns natural language prompts into coding suggestions across dozens of languages. These features make Copilot ideal for developing applications, writing unit tests, and authoring documentation. You can use GitHub Copilot to assist with writing software applications in nearly any popular language, including Go.
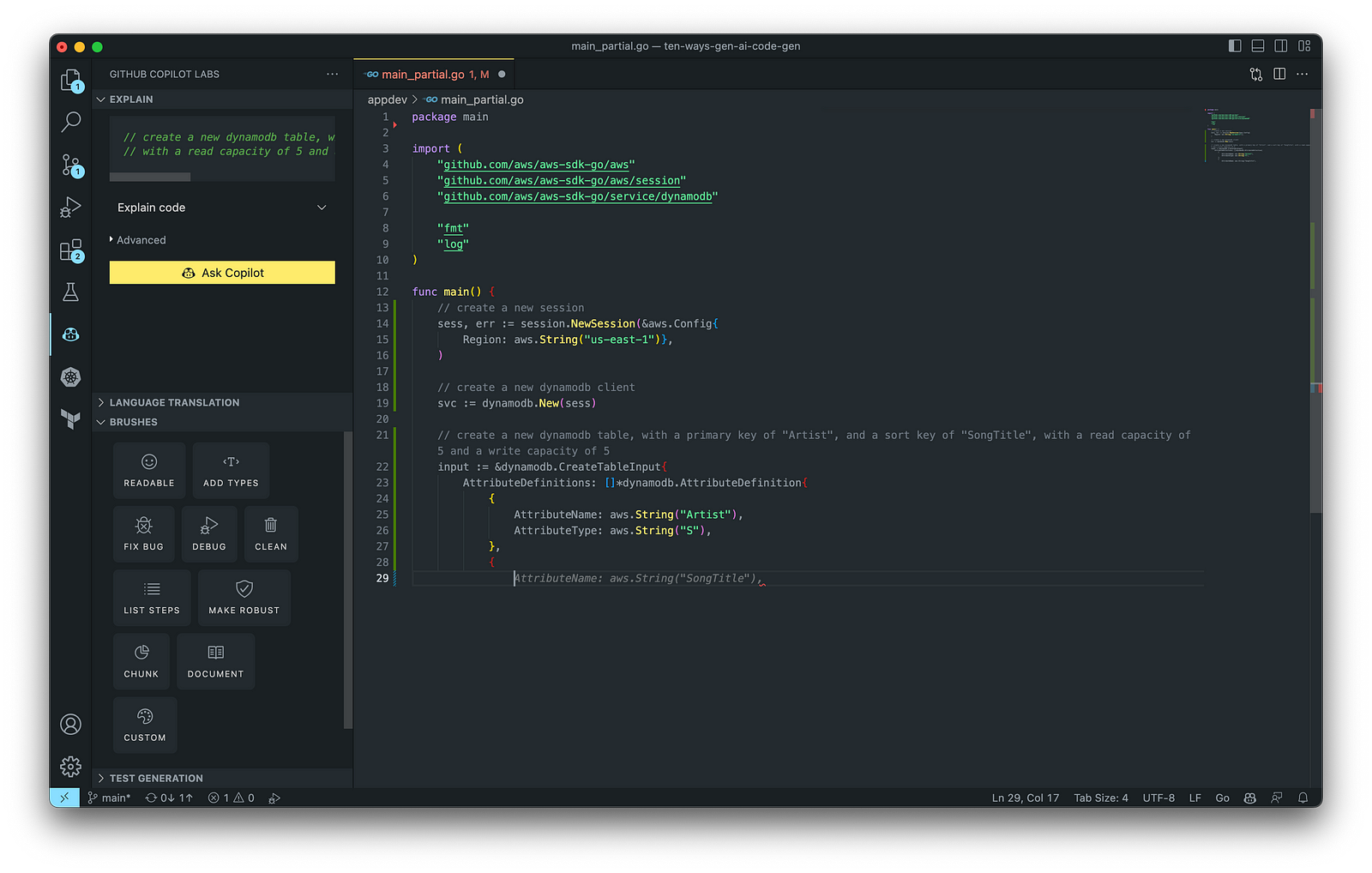
The final application, which uses the AWS SDK for Go to create an Amazon DynamoDB table, shown below, was formatted using the Go extension by Google and optimized using the ‘Readable,’ ‘Make Robust,’ and ‘Fix Bug’ GitHub Code Brushes.
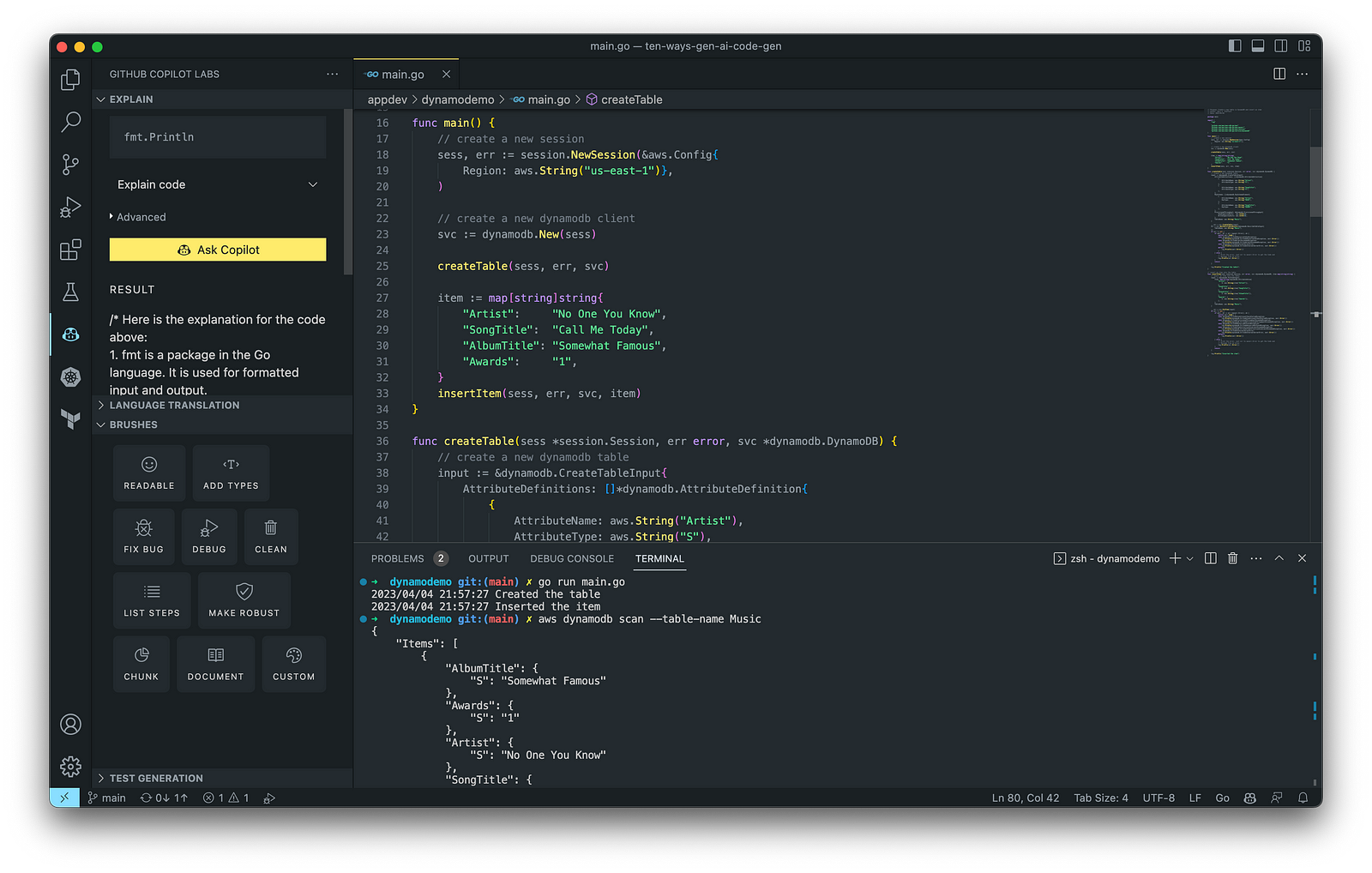
Generating Unit Tests
Using JavaScript and TypeScript, you can take advantage of TestPilot to generate unit tests based on your existing code and documentation. TestPilot, part of GitHub Copilot Labs, uses GitHub Copilot’s AI technology.

2. Infrastructure as Code (IaC)
Widespread Infrastructure as Code (IaC) tools include Pulumi, AWS CloudFormation, Azure ARM Templates, Google Deployment Manager, AWS Cloud Development Kit (AWS CDK), Microsoft Bicep, and Ansible. Many IaC tools, except AWS CDK, use JSON- or YAML-based domain-specific languages (DSLs).
AWS CloudFormation
AWS CloudFormation is an Infrastructure as Code (IaC) service that allows you to easily model, provision, and manage AWS and third-party resources. The CloudFormation template is a JSON or YAML formatted text file. You can use GitHub Copilot to assist with writing IaC, including AWS CloudFormation in either JSON or YAML.

You can use the YAML Language Support by Red Hat extension to write YAML in VS Code.

VS Code has native JSON support with JSON Schema Store, which includes AWS CloudFormation. VS Code uses the CloudFormation schema for IntelliSense and flag schema errors in templates.

HashiCorp Terraform
In addition to AWS CloudFormation, HashiCorp Terraform is an extremely popular IaC tool. According to HashiCorp, Terraform lets you define resources and infrastructure in human-readable, declarative configuration files and manages your infrastructure’s lifecycle. Using Terraform has several advantages over manually managing your infrastructure.
Terraform plugins called providers let Terraform interact with cloud platforms and other services via their application programming interfaces (APIs). You can use the AWS Provider to interact with the many resources supported by AWS.
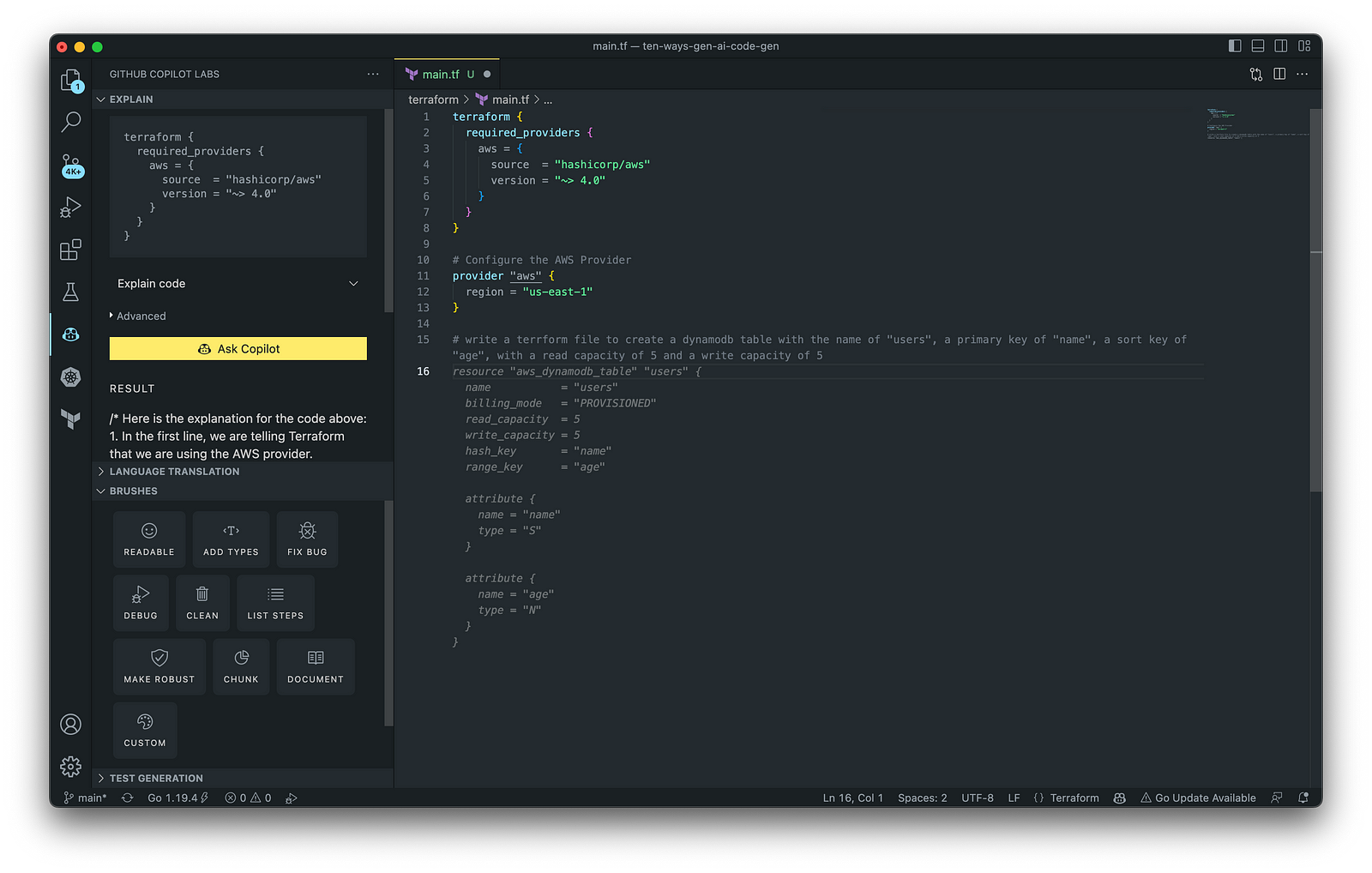
3. AWS Lambda
Lambda, according to AWS, is a serverless, event-driven compute service that lets you run code for virtually any application or backend service without provisioning or managing servers. You can trigger Lambda from over 200 AWS services and software as a service (SaaS) applications and only pay for what you use. AWS Lambda natively supports Java, Go, PowerShell, Node.js, C#, Python, and Ruby. AWS Lambda also provides a Runtime API allowing you to use additional programming languages to author your functions.
You can use GitHub Copilot to assist with writing AWS Lambda functions in any of the natively supported languages. You can further optimize the resulting Lambda code with GitHub’s Code Brushes.

The final Python-based AWS Lambda, below, was formatted using the Black Formatter and Flake8 extensions and optimized using the ‘Readable,’ ‘Debug,’ ‘Make Robust,’ and ‘Fix Bug’ GitHub Code Brushes.

You can easily convert the Python-based AWS Lambda to Java using GitHub Copilot Lab’s ability to translate code between languages. Install the GitHub Copilot Labs extension for VS Code to try out language translation.

4. IAM Policies
AWS Identity and Access Management (AWS IAM) is a web service that helps you securely control access to AWS resources. According to AWS, you manage access in AWS by creating policies and attaching them to IAM identities (users, groups of users, or roles) or AWS resources. A policy is an object in AWS that defines its permissions when associated with an identity or resource. IAM policies are stored on AWS as JSON documents. You can use GitHub Copilot to assist in writing IAM Policies.
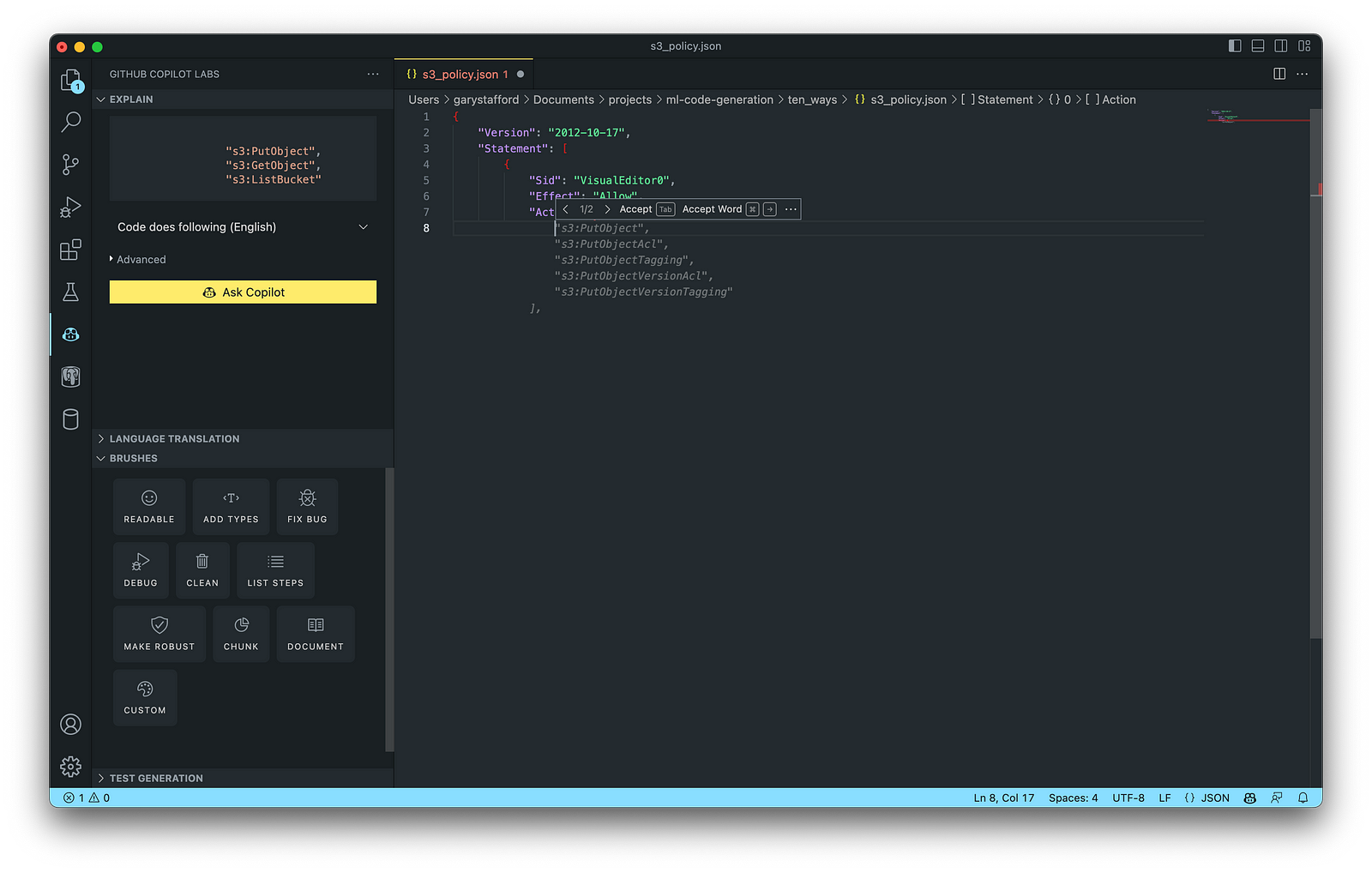
The final AWS IAM Policy, below, was formatted using VS Code’s built-in JSON support.

5. Structured Query Language (SQL)
SQL has many use cases on AWS, including Amazon Relational Database Service (RDS) for MySQL, PostgreSQL, MariaDB, Oracle, and SQL Server databases. SQL is also used with Amazon Aurora, Amazon Redshift, Amazon Athena, Apache Presto, Trino (PrestoSQL), and Apache Hive on Amazon EMR.
You can use IDEs like VS Code with its SQL dialect-specific language support and formatted extensions. You can further optimize the resulting SQL statements with GitHub’s Code Brushes.

The final PostgreSQL script, below, was formatted using the Sql Formatter extension and optimized using the ‘Readable’ and ‘Fix Bug’ GitHub Code Brushes.

6. Big Data
Big Data, according to AWS, can be described in terms of data management challenges that — due to increasing volume, velocity, and variety of data — cannot be solved with traditional databases. AWS offers managed versions of Apache Spark, Apache Flink, Apache Zepplin, and Jupyter Notebooks on Amazon EMR, AWS Glue, and Amazon Kinesis Data Analytics (KDA).
Apache Spark
According to their website, Apache Spark is a multi-language engine for executing data engineering, data science, and machine learning on single-node machines or clusters. Spark jobs can be written in various languages, including Python (PySpark), SQL, Scala, Java, and R. Apache Spark is available on a growing number of AWS services, including Amazon EMR and AWS Glue.

The final Python-based Apache Spark job, below, was formatted using the Black Formatter extension and optimized using the ‘Readable,’ ‘Document,’ ‘Make Robust,’ and ‘Fix Bug’ GitHub Code Brushes.

7. Configuration and Properties Files
According to TechTarget, a configuration file (aka config) defines the parameters, options, settings, and preferences applied to operating systems, infrastructure devices, and applications. There are many examples of configuration and properties files on AWS, including Amazon MSK Connect (Kafka Connect Source/Sink Connectors), Amazon OpenSearch (Filebeat, Logstash), and Amazon EMR (Apache Log4j, Hive, and Spark).
Kafka Connect
Kafka Connect is a tool for scalably and reliably streaming data between Apache Kafka and other systems. It makes it simple to quickly define connectors that move large collections of data into and out of Kafka. AWS offers a fully-managed version of Kafka Connect: Amazon MSK Connect. You can use GitHub Copilot to write Kafka Connect Source and Sink Connectors with Kafka Connect and Amazon MSK Connect.

The final Kafka Connect Source Connector, below, was formatted using VS Code’s built-in JSON support. It incorporates the Debezium connector for MySQL, Avro file format, schema registry, and message transformation. Debezium is a popular open source distributed platform for performing change data capture (CDC) with Kafka Connect.

8. Apache Airflow DAGs
Apache Airflow is an open-source platform for developing, scheduling, and monitoring batch-oriented workflows. Airflow’s extensible Python framework enables you to build workflows connecting with virtually any technology. DAG (Directed Acyclic Graph) is the core concept of Airflow, collecting Tasks together, organized with dependencies and relationships to say how they should run.
Amazon Managed Workflows for Apache Airflow (Amazon MWAA) is a managed orchestration service for Apache Airflow. You can use GitHub Copilot to assist in writing DAGs for Apache Airflow, to be used with Amazon MWAA.

The final Python-based Apache Spark job, below, was formatted using the Black Formatter extension. Unfortunately, based on my testing, code optimization with GitHub’s Code Brushes is impossible with Airflow DAGs.

9. Containerization
According to Check Point Software, Containerization is a type of virtualization in which all the components of an application are bundled into a single container image and can be run in isolated user space on the same shared operating system. Containers are lightweight, portable, and highly conducive to automation. AWS describes containerization as a software deployment process that bundles an application’s code with all the files and libraries it needs to run on any infrastructure.
AWS has several container services, including Amazon Elastic Container Service (Amazon ECS), Amazon Elastic Kubernetes Service (Amazon EKS), Amazon Elastic Container Registry (Amazon ECR), and AWS Fargate. Several code-based resources can benefit from a Generative AI coding tool like GitHub Copilot, including Dockerfiles, Kubernetes resources, Helm Charts, Weaveworks Flux, and ArgoCD configuration.
Kubernetes
Kubernetes objects are represented in the Kubernetes API and expressed in YAML format. Below is a Kubernetes Deployment resource file, which creates a ReplicaSet to bring up multiple replicas of nginx Pods.

The final Kubernetes resource file below contains Deployment and Service resources. In addition to GitHub Copilot, you can use Microsoft’s Kubernetes extension for VS Code to use IntelliSense and flag schema errors in the file.
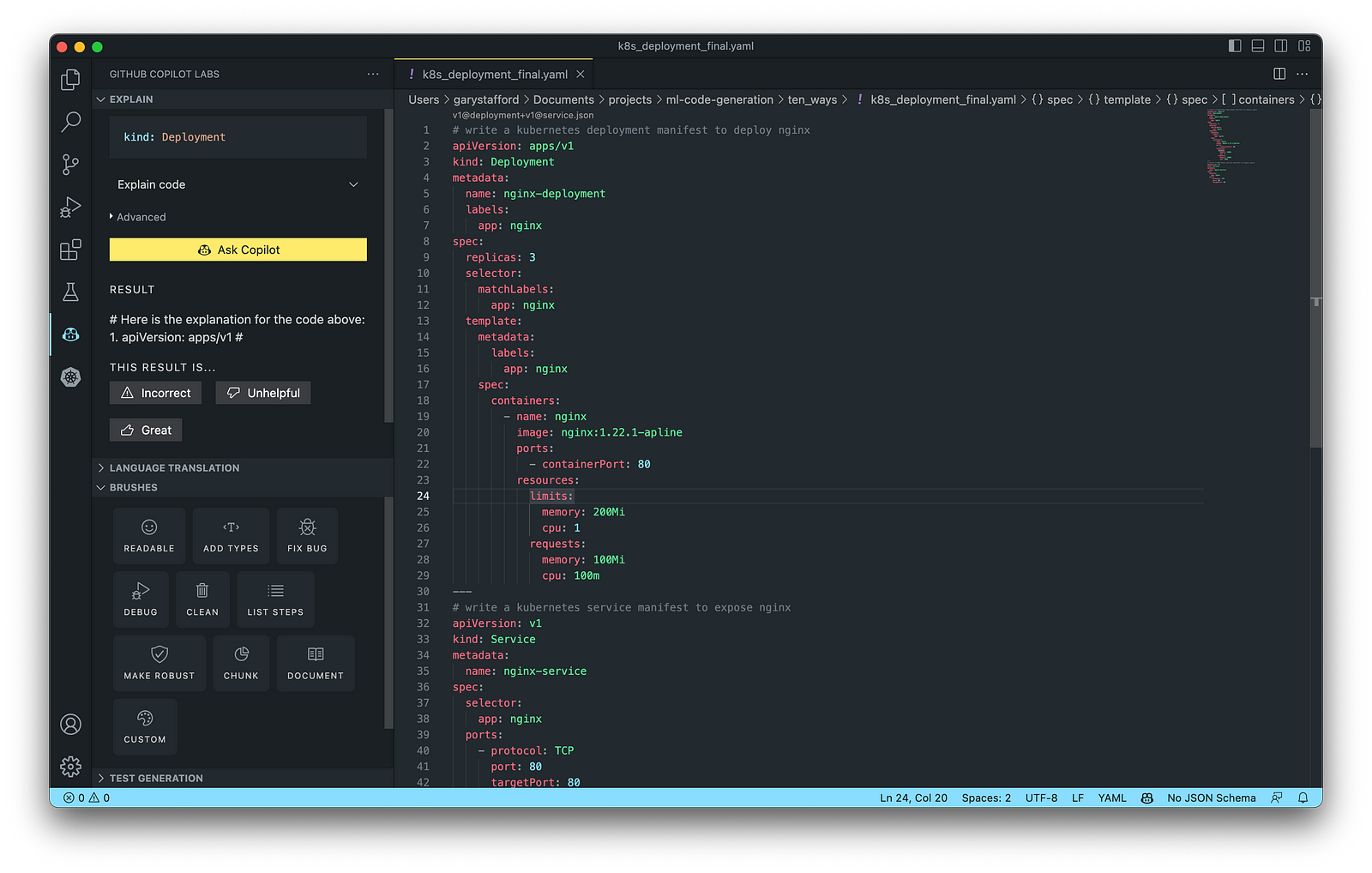
10. Utility Scripts
According to Bing AI — Search, utility scripts are small, simple snippets of code written as independent code files designed to perform a particular task. Utility scripts are commonly written in Bash, Shell, Python, Ruby, PowerShell, and PHP.
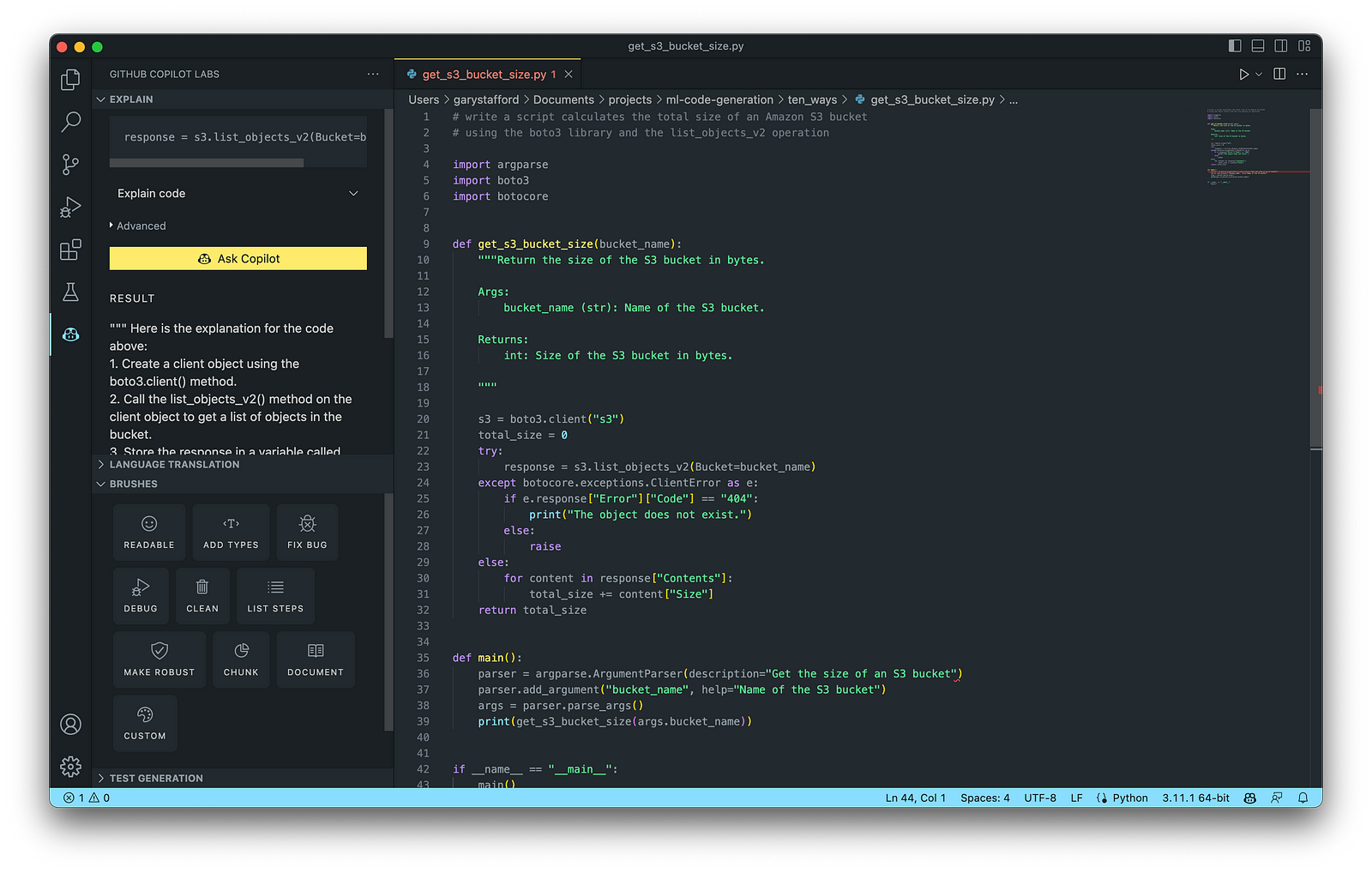
AWS utility scripts leverage the AWS Command Line Interface (AWS CLI) for Bash and Shell and AWS SDK for other programming languages. SDKs take the complexity out of coding by providing language-specific APIs for AWS services. For example, Boto3, AWS’s Python SDK, easily integrates your Python application, library, or script with AWS services, including Amazon S3, Amazon EC2, Amazon DynamoDB, and more.

An example of a Python script to calculate the total size of an Amazon S3 bucket, below, was inspired by 100daysofdevops/N-days-of-automation, a fantastic set of open source AWS-oriented automation scripts.
Conclusion
In this post, you learned ten ways to leverage Generative AI coding tools like GitHub Copilot for development on AWS. You saw how combining the latest generation of Generative AI coding tools, a mature and extensible IDE, and your coding experience will accelerate development, increase productivity, and reduce cost.
🔔 To keep up with future content, follow Gary Stafford on LinkedIn.
This blog represents my viewpoints and not those of my employer, Amazon Web Services (AWS). All product names, logos, and brands are the property of their respective owners.
Accelerate Software Development with Six Popular Generative AI-Powered Coding Tools
Posted by Gary A. Stafford in AI/ML, Cloud, Software Development on March 25, 2023
Explore six popular generative AI-powered tools, including ChatGPT, Copilot, CodeWhisperer, Tabnine, Bing, and ChatSonic

Introduction
Modern software systems continue to grow inherently more complex over time. We have evolved from bulky monoliths to loosely coupled, event-driven, fault-tolerant, stateless, serverless, cloud-native, real-time, microservices-based, API-first, continuously deployed distributed systems, festooned with CQRS, 2PC, DDD, EDA, DOMA, Sagas, BFFs, GraphQL, gRPC, micro-frontends, contract tests, and Hexagonal architectures. Generative AI-powered coding tools do not necessarily make a developer’s job easier — they assist developers in dealing with increasing system complexity.
This post examines six popular generative AI-powered coding tools, including chat-based OpenAI ChatGPT, Microsoft’s all-new Bing Chat, and ChatSonic, as well as IDE-based Tabnine, GitHub Copilot, and Amazon CodeWhisperer (Preview). In this post, each tool will assist with developing an identical program to complete a series of common tasks on AWS. We will then compare and contrast each tool’s ease of use and the resulting code accuracy and quality.
“Generative AI coding tools are a new class of software development tools that leverage machine learning algorithms to assist developers in writing code. These tools use AI models trained on vast amounts of code to offer suggestions for completing code snippets, writing functions, and even entire blocks of code.” (quote generated by ChatGPT)
The generative AI space is evolving at a breakneck pace. Tools continue to rapidly improve their AI models, add new features, and adjust pricing. In just the short time it took to research and write this article:
- OpenAI announced GPT-4 on March 14, 2023
- Microsoft announced new Bing running on GPT-4 on March 14, 2023
- Google’s Bard AI launched in Early Access on March 21, 2023
- GPT-4 was added to Azure’s OpenAI Service on March 21, 2023
- Writesonic ChatSonic revised its pricing model on March 21, 2023
- GitHub announced GitHub Copilot X on March 22, 2023
- OpenAI announced ChatGPT plugins on March 23, 2023
Generative AI
According to McKinsey & Company in their recent article, What is generative AI?, “Generative artificial intelligence (AI) describes algorithms (such as ChatGPT) that can be used to create new content, including audio, code, images, text, simulations, and videos. Recent new breakthroughs in the field have the potential to drastically change the way we approach content creation.”
Generative AI for Code Generation
According to Papers with Code, “Code Generation is an important field to predict explicit code or program structure from multimodal data sources such as incomplete code, programs in another programming language, natural language descriptions or execution examples. Code Generation tools can assist the development of automatic programming tools to improve programming productivity.” Similarly, according to MarketTechPost, “Generative AI technologies have led to a surge of interest and progress in code generation applications. These technologies use machine learning algorithms and natural language processing to assist developers in automating the time-consuming and laborious portions of coding.”
Common Features
Standard features of leading generative AI-powered coding tools include the following:
- Whole-line, full-function, and block code completion
- Natural language to code completion
- Code suggestions based on the model’s pre-trained dataset
- Context-aware recommendations based on your existing code
- Context-aware recommendations based on your code comments
- Native integration with popular IDEs
- Multi-language coding support
- Respond to follow-up instructions (resulting in refinement of code)
Benefits
The benefits of generative AI-powered code generation include the following:
- Accelerate application development
- Improve development productivity
- Decrease development costs
- Produce higher-quality, more consistent code
- Enable better code documentation and test coverage
- Learn new programming languages and coding techniques
- Contributes to the democratization of programming
Methods of Code Generation
As you explore different generative AI tools for writing code, you will likely develop techniques for generating effective code based on the tool.
Long-form Narrative Approach
With chat-based tools like OpenAI ChatGPT, Microsoft Bing Chat, and ChatSonic, developers might use a longer narrative-type instruction (aka one-shot approach) rather than a series of more concise instructions to generate the code. For example:
- “Write a Python script to create a new DynamoDB table with a partition key of username, a sort key of last_name, and provisioned throughput of 5 RCUs and 5 WCUs. Pass the table name into the function as a variable. Wait until the table exists before continuing. Include try/except blocks and logging. If the table fails to be created, log a FATAL error and exit, and if the table already exists, log a WARNING. Add a main function and wrap everything in a Class called DynamoUtilities.”
- “Add three new functions, including a function to insert a new item into a table, a function to retrieve an item from a table, and a function to delete a table.”
Below is an example of OpenAI ChatGPT’s response using this approach. Of course, this method doesn’t limit you from following up with additional instructions to further enhance and refine the code.

Progressive Approach
Although the previous approach makes for an impressive demonstration, that method of writing code differs from how most developers work. Instead, a developer might use a progressive approach (aka chain of thought) to generate and refine the code with chat-based tools. In my tests with chat-based tools, I found a progressive approach of asking multiple, concise instructions to guide the model toward the desired behavior produced higher-quality code with fewer errors. For example:
- “Write a Python script to create a DynamoDB table with a partition key of username, a sort key of last_name, and provisioned throughput of 5 RCUs and 5 WCUs.”
- “Pass the table name into the create table function as a variable.”
- “Wait until the table exists before continuing.”
- “Add logging with the default level of INFO.”
- “Include try/except blocks, log a FATAL error and exit if the table fails to be created and a WARNING if the table already exists.”
- “Add a function to delete the table and wait until the table is deleted before continuing.”
- “Add a main function.”
- “Add a function to insert a new item into a table and a function to get an item from a table.”
- “Wrap the code in a Class called DynamoUtilities.”
- “Convert the Python code to Java.”
Below is an example of OpenAI ChatGPT’s response using a progressive code generation approach. The final code results from an initial prompt and a series of follow-up instructions to enhance and refine the code.
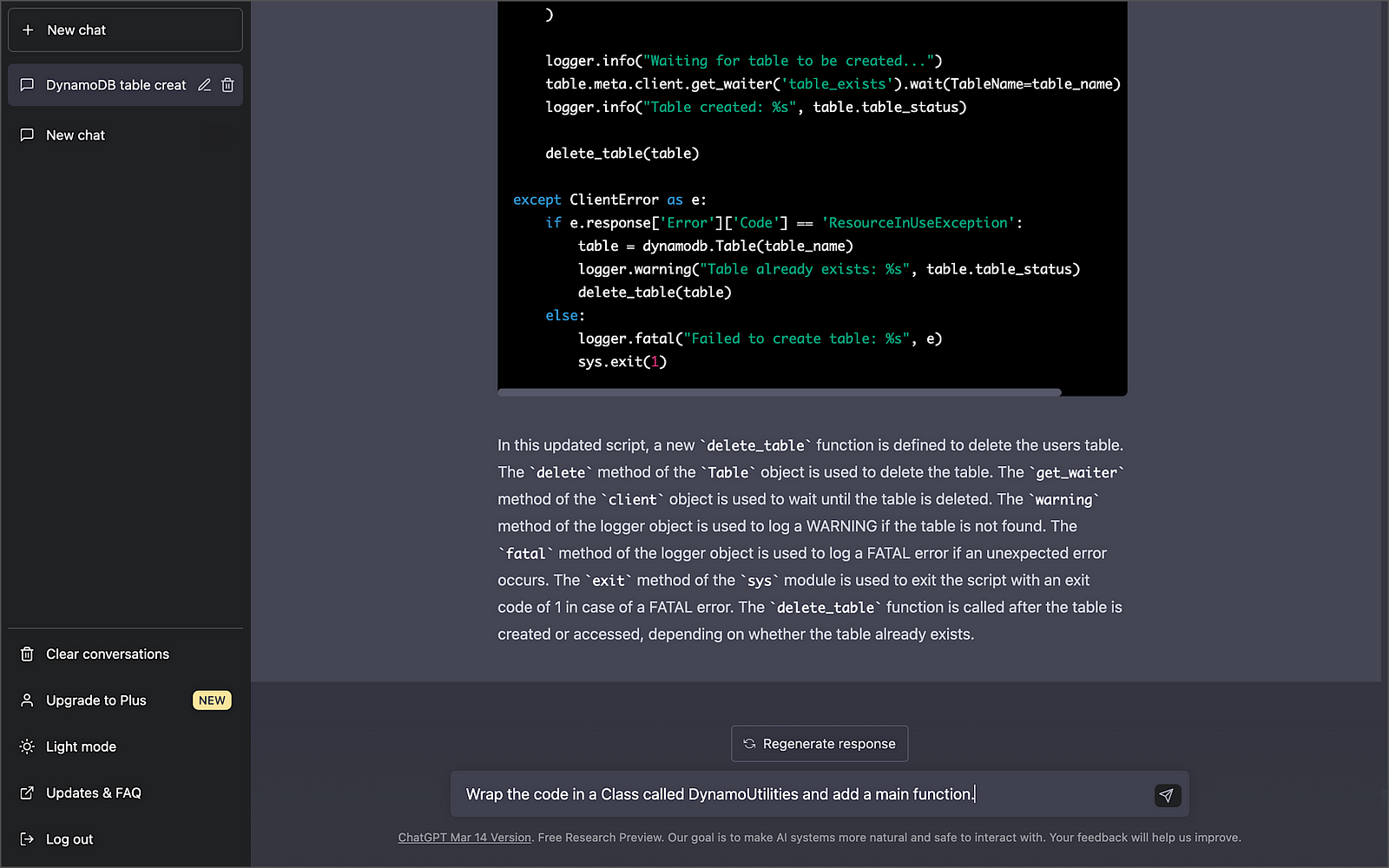
Be aware that most chat-based tools have a session limit. For example, Bing has a limit of 15 chats per session, which will limit the number of follow-up instructions you can use to modify the code. Additionally, the slightest variation in how an instruction is phrased may significantly impact the code generated. There is a whole science and blossoming industry around prompt optimization!
IDE-based Code Generation
Tabnine, GitHub Copilot, and Amazon CodeWhisperer use generative AI technology to predict and suggest new lines or blocks of code based on their trained pre-model and the context and syntax of existing code and code comments within your IDE. These tools differ from tools like OpenAI ChatGPT, Microsoft Bing Chat, and ChatSonic, which can generate code using a chat-based interaction with the user. IDE-based tools like Tabnine, GitHub Copilot, and Amazon CodeWhisperer are like having an AI-powered paired programming partner that enhances your development productivity. However, these tools also require a reasonable level of development experience, in my opinion.
Below, we see an example of Tabnine Pro’s ability to generate whole-line code completion (line 11) and full-function code completion (lines 12–41) based on the existing code context and syntax (lines 1–11).
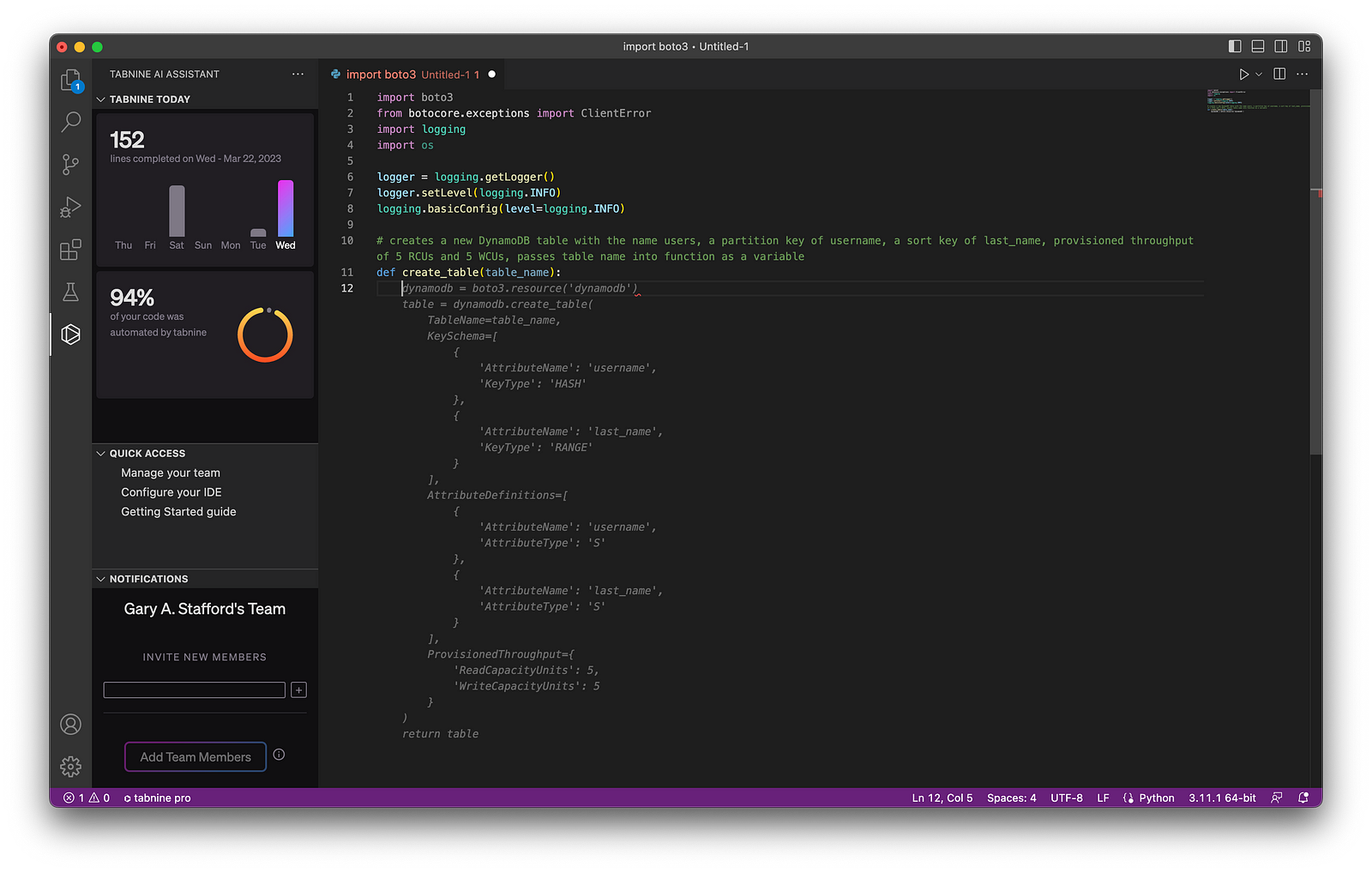
Although writing code comments may be perceived as easier than using generative code completion, many industry pundits dissuade what they call “ comment-driven development.” Instead, they believe using generative AI-based code completion results in superior results.
Testing Generative AI Tools
With the assistance of each tool, I have written a Python script to perform the following tasks: 1) create an Amazon DynamoDB table, 2) insert an item into a table, 3) retrieve an item from a table, and finally, 4) delete a table. Although I tested the results in multiple IDEs, I only included the results from Visual Studio Code (VS Code). Tool interactions and code results were consistent across IDEs.
As a reference for “what good looks like” when evaluating code accuracy and quality, I referenced the examples provided with AWS’s Boto3 SDK and DynamoDB API documentation and the PEP 8 — Style Guide for Python Code.

An example of the final code, written with the assistance of GitHub Copilot, including unit tests, is available on GitHub.
OpenAI ChatGPT

According to Wikipedia, ChatGPT (Generative Pre-trained Transformer) “is an artificial intelligence chatbot developed by OpenAI and launched in November 2022. It is built on top of OpenAI GPT-3 and GPT-4 [released March 14, 2023] families of large language models [LLMs] and has been fine-tuned (an approach to transfer learning) using both supervised and reinforcement learning techniques.”
According to the VC-backed California-based startup OpenAI, “ChatGPT interacts in a conversational way. The dialogue format makes it possible for ChatGPT to answer follow-up questions, admit its mistakes, challenge incorrect premises, and reject inappropriate requests.” Further, “ChatGPT was optimized for dialogue using Reinforcement Learning with Human Feedback (RLHF), which uses human demonstrations and preference comparisons to guide the model toward desired behavior.”
Getting Started with ChatGPT
You can get started with ChatGPT for free using the Free Plan. However, if you want to use the latest generative AI models, get faster results, and ensure availability, you should consider ChatGPT Plus for $20/month. GPT-4 is now available through the ChatGPT Plus Plan.

ChatGPT’s Free Plan is a great way to test the generative AI waters. However, due to potentially slow chat responses and unavailability during peak times, you will want to upgrade to ChatGPT Plus if your team plans to rely on the tool.

Using ChatGPT
OpenAI ChatGPT is available through a web-based chat interface or programmatically using OpenAI APIs. For API users, OpenAI provides online API references, code examples, and an interactive coding playground. According to OpenAI, “The OpenAI API can be applied to virtually any task that involves understanding or generating natural language, code, or images.”

Below is an example of output using ChatGPT’s UI. Using the UI, ChatGPT’s results are a mix of code blocks and explanatory text. ChatGPT has a straightforward, clean, and functional web-based user interface. There is a button to copy the resulting code to your clipboard, and previous chat sessions are saved and accessible.
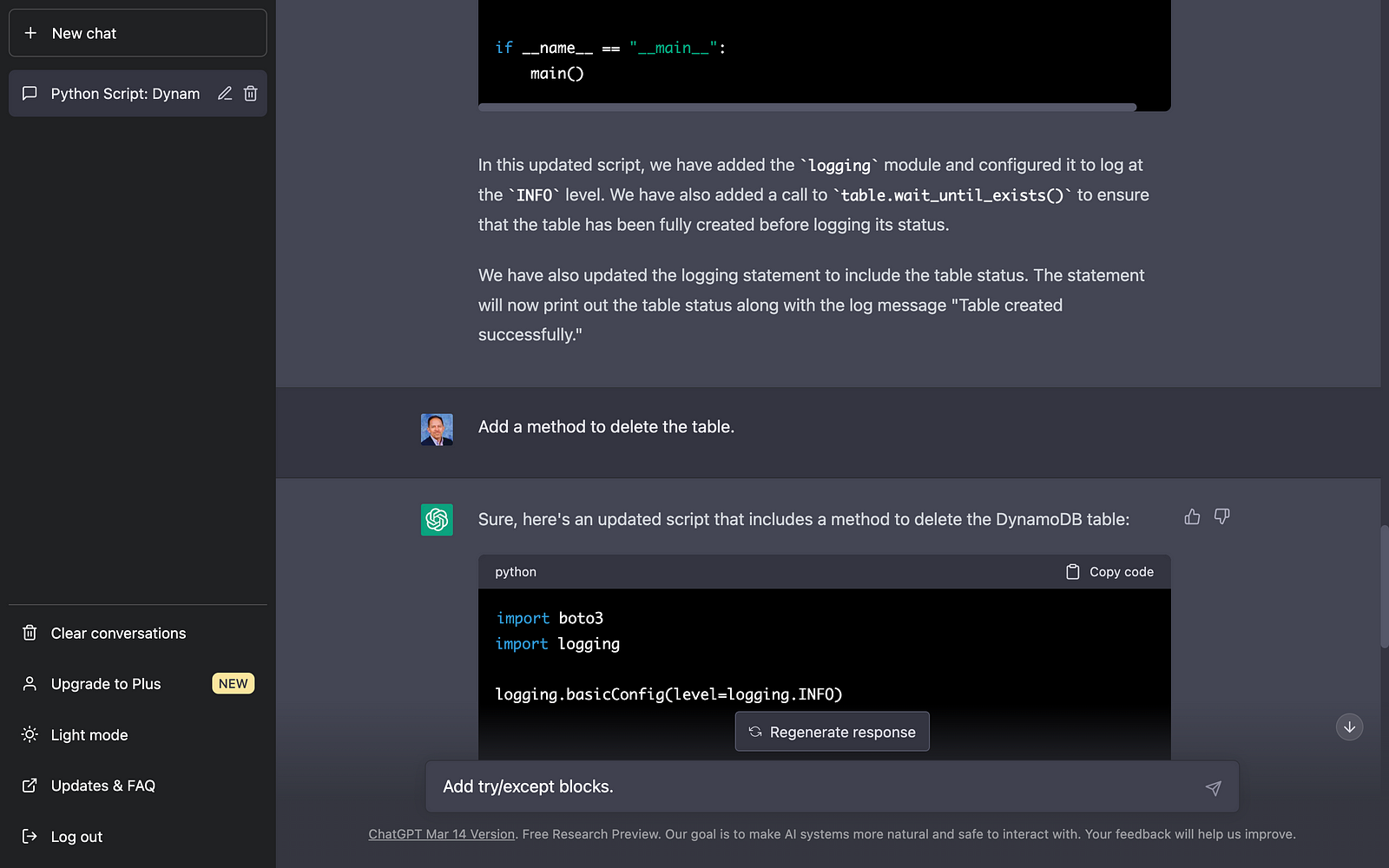
ChatGPT Results
In my tests, using a progressive chat approach to building the code with concise instructional questions versus a longer-form narrative approach produced better-quality code. The code created using a progressive chat approach was accurate, error-free, and well-written.
A significant advantage of ChatGPT is its ability to remember what happened earlier in the conversation. According to OpenAI documentation, the model can reference up to approximately 3k words (or 4k tokens) from the current conversation; any information beyond that is not stored.

Microsoft Bing Chat
According to their blog post on February 7, Microsoft launched “an all new, AI-powered Bing search engine and Edge browser, available in preview now at Bing.com, to deliver better search, more complete answers, a new chat experience and the ability to generate content. We think of these tools as an AI copilot for the web.” The latest version of Bing “is running on a new, next-generation OpenAI large language model that is more powerful than ChatGPT and customized specifically for search. It takes key learnings and advancements from ChatGPT and GPT-3.5 — and it is even faster, more accurate and more capable.” On March 14, Microsoft confirmed the new Bing runs on OpenAI’s latest GPT-4.
Getting Started with Bing Chat
Bing’s Chat mode is only available when you access Bing using Microsoft Edge.

Microsoft Edge is available for most major platforms, including MacOS, iOS, and Android.

Using Bing Chat
Bing Chat now supports up to 15 chats per session and 150 per day. Users can supply up to 15 prompts (questions or instructions) to create and improve the code results. When responding, Bing considers the context of previous prompts in the same chat session.

Bing Chat Results
Like OpenAI ChatGPT, Bing Chat delivered accurate, error-free, and well-written code. Although ChatGPT and Bing Chat are not IDE-based tools, the resulting code is easily copied into your project or saved to disk. Additionally, these tools provide an excellent opportunity to learn new languages and coding techniques without requiring an IDE.

ChatSonic

ChatSonic is a service of Writesonic. According to Y Combinator, startup Writesonic, founded by Samanyou Garg, is backed by leading venture capital firms, including Y Combinator, HOF Capital, Rebel Fund, Soma Capital, Broom Ventures, Amino Capital, and some of the best angels from different industries.
ChatSonic claims to improve upon the limitations of ChatGPT. According to Wordsonic, ChatSonic is “a powerful chatbot, powered by Google Search, allowing it to provide up-to-date and accurate content and hence superior to ChatGPT, which can not generate content on topics after September 2021. Additionally, it can create digital images and respond to voice commands and many more additional features. So, ChatSonic has addressed all the limitations of ChatGPT.”
Using ChatSonic
You can get started with ChatSonic for free, with up to 10,000 words. However, like OpenAI ChatGPT, if you want to take advantage of GPT-4, you will want to upgrade to a paid plan, starting at $13/month with an annual commitment.

Using ChatSonic is nearly identical to using OpenAI ChatGPT.

During my testing for this article, I ran into several issues with ChatSonic’s Premium Plan’s Free Trial. For example, the code block constantly flashed and refreshed on the screen using Chrome for Mac; a minor issue but highly annoying. Also, when copying or downloading code results, you get both code and the informational text, which you must separate. Worse, the final code block did not always include all the code output; the code was intermixed with informational text. Lastly, when using follow-up instructions to modify the results, the regenerated code often missed previous code modifications. In my tests, after several follow-up instructions, the resulting code was missing up to one-third of the previous statements. Due to these issues, the resulting code block was malformed and unrunnable. The more I chatted with ChatSonic, the worse the code issues became, and the issues were repeatable.


ChatSonic Results
Unlike ChatGPT and Bing Chat, ChatSonic’s Premium Plan’s Free Trial did not deliver accurate, error-free, or well-written code during my tests. Although later tests with fewer follow-up instructions provided better though incomplete results, my first impressions were unfavorable. The results might be improved using ChatSonic’s Superior or Ultimate Plans, which use the newest GPT-4 model. However, given the results of Premium, I was not keen to invest in the paid plan to find out.
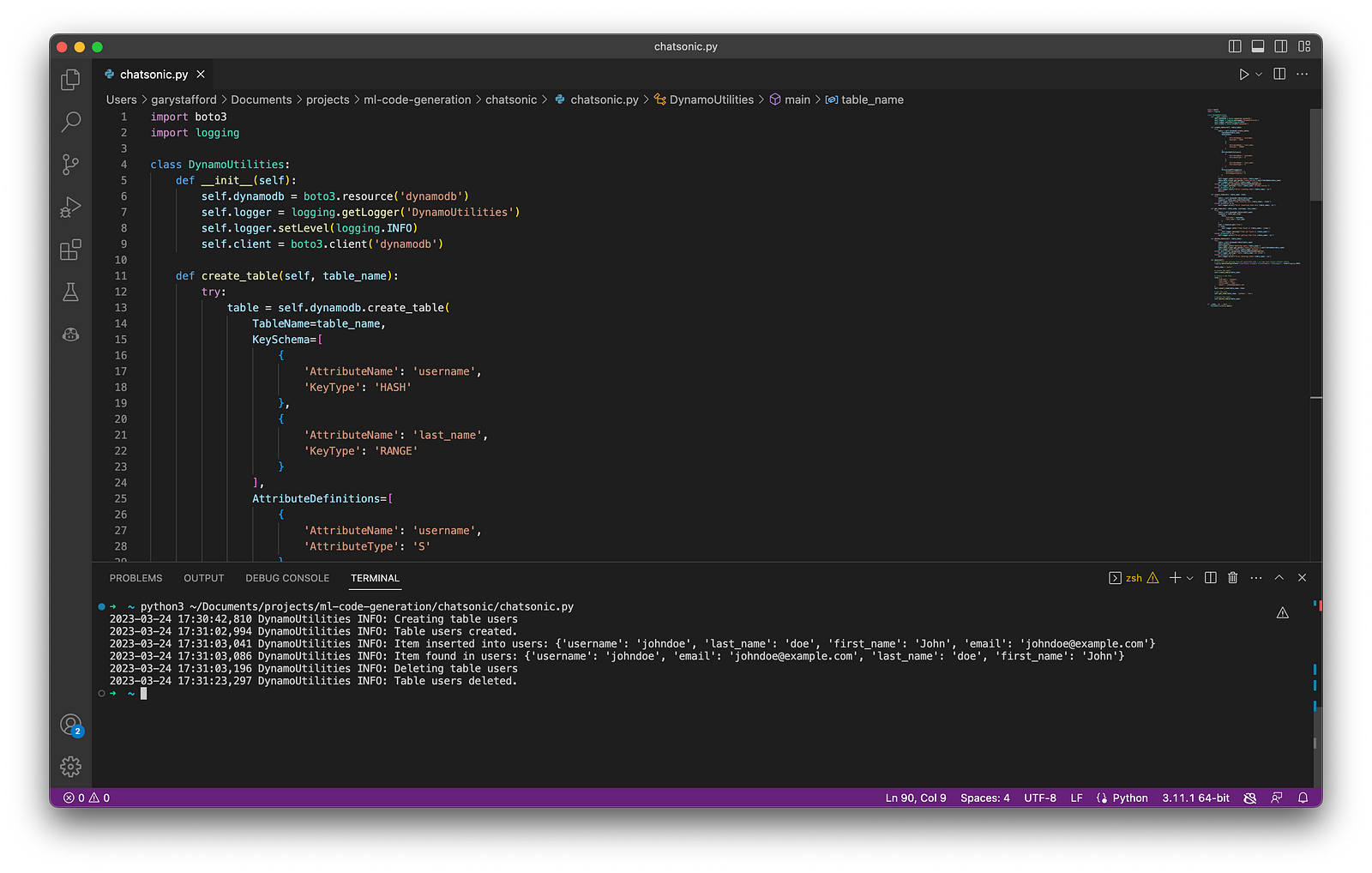
Tabnine

According to the VC-backed, Israel-based startup Tabnine, “Tabnine is to create and deliver a top-to-bottom AI-assisted development workflow that empowers all code creators, in all languages, from concept through to completion.” Tabnine Pro serves whole-line, full-function, and natural language to code completions. Regarding AI, in Tabline’s opinion, “a multi-model approach far outperforms a monolithic approach. That’s why we’ve developed language-specific code native AI models, which are pre-trained on code and provide faster and more accurate code completions based on your tech stack.” To learn more about their models, read Tabnine’s blog post, Tabnine Enterprise vs. ChatGPT Plus.
Getting Started with Tabline
You can get started with Tabline for free with the Starter plan. However, the features of the Starter plan are limited compared to the Pro Plan, which starts at $12/month for a single user.

You can try out the Pro plan’s features, which are free for 14 days.

Tabnine supports a wide range of programming languages.
To get started with Tabline, choose your favorite IDE and install the required extensions or plugins.

Tabnine provides automated or manual IDE installation.

The Tabnine extension for VS Code already has a staggering 5.1 million downloads! A valuable tip learned from testing the IDE-based tools was to disable all non-essential extensions to give me a baseline of the tool’s capabilities. Initially, with over eighty VS Code extensions enabled, I found it hard to separate and understand each tool’s features and potential conflicts with other extensions I had loaded, especially other generative AI tools.

Using Tabnine
As discussed earlier, Tabnine, like GitHub Copilot and Amazon CodeWhisperer, uses generative AI technology to predict and suggest subsequent lines of code based on context and syntax within your IDE. It is like having IntelliSense on steroids.
Like Copilot and CodeWhisperer, it takes some practice to get efficient with Tabnine. After starting with the initial import and logging statements, I used code comments to generate the functions, then returned to add exception-catching and logging.
Tabnine, like Copilot and CodeWhisperer, includes a collapsible navigation bar. However, unlike those tools, I found Tabnine’s navigation bar provided little value, in my opinion. It seemed more like marketing stats and a way to encourage more users to sign-up rather than providing useful development features like Copilot and CodeWhisperer.
Below, we see an example of Tabnine Pro’s ability to generate whole-line code completion (line 11) and full-function code completion (lines 12–41) based on the existing code context and syntax (lines 1–11).

Tabnine Results
Given my development knowledge and experience with similar tools, I found the final code results with Tabnine Pro were accurate, error-free, and well-formed (Pythonic). However, I would have liked to see additional features to differentiate Tabnine from its competitors.

GitHub Copilot

According to GitHub, “Copilot is an AI pair programmer that offers autocomplete-style suggestions as you code. You can receive suggestions from GitHub Copilot either by starting to write the code you want to use or by writing a natural language comment describing what you want the code to do. In addition, GitHub Copilot analyzes the context in the file you are editing and related files, and offers suggestions from within your text editor.” Copilot is powered by OpenAI Codex, a new AI system created by OpenAI.
Given the vast popularity and volume of code on GitHub, it is no wonder Copilot is trained in all languages that appear in GitHub’s public repositories. GitHub points out that the quality of suggestions you receive may depend on the volume and diversity of training data for that language.
GitHub Copilot X
GitHub is extending Copilot’s capabilities, with the announcement of GitHub Copolit X on March 21, 2023. According to GitHub, “With chat and terminal interfaces, support for pull requests, and early adoption of OpenAI’s GPT-4, GitHub Copilot X is our vision for the future of AI-powered software development. Integrated into every part of your workflow.”
Getting Started with Copilot
Get started with GitHub Copilot for as little as $10/month. This Monthly Plan is a great way to test Copilot’s capabilities. In addition, GitHub is currently offering a 60-day free trial of Copilot.

Regarding IDE support, Copilot is currently available as an extension in Visual Studio Code, Visual Studio, Neovim, and the JetBrains suite of IDEs. The GitHub Copilot extension already has 4.2 million downloads, and the GitHub Copilot Nightly extension, used for this post, has almost 250,000 downloads.

GitHub Copilot can also be used with Codespaces, GitHub’s fully configured development environments in the Cloud based on VS Code, similar to AWS Cloud9.
Using Copilot
As discussed earlier, GitHub Copilot, like Tabnine and Amazon CodeWhisperer, uses generative AI technology to predict and suggest new lines of code based on existing context and syntax. In addition, Copilot includes an inline pop-up toolbar, which makes it easier to scroll through and accept code choices.
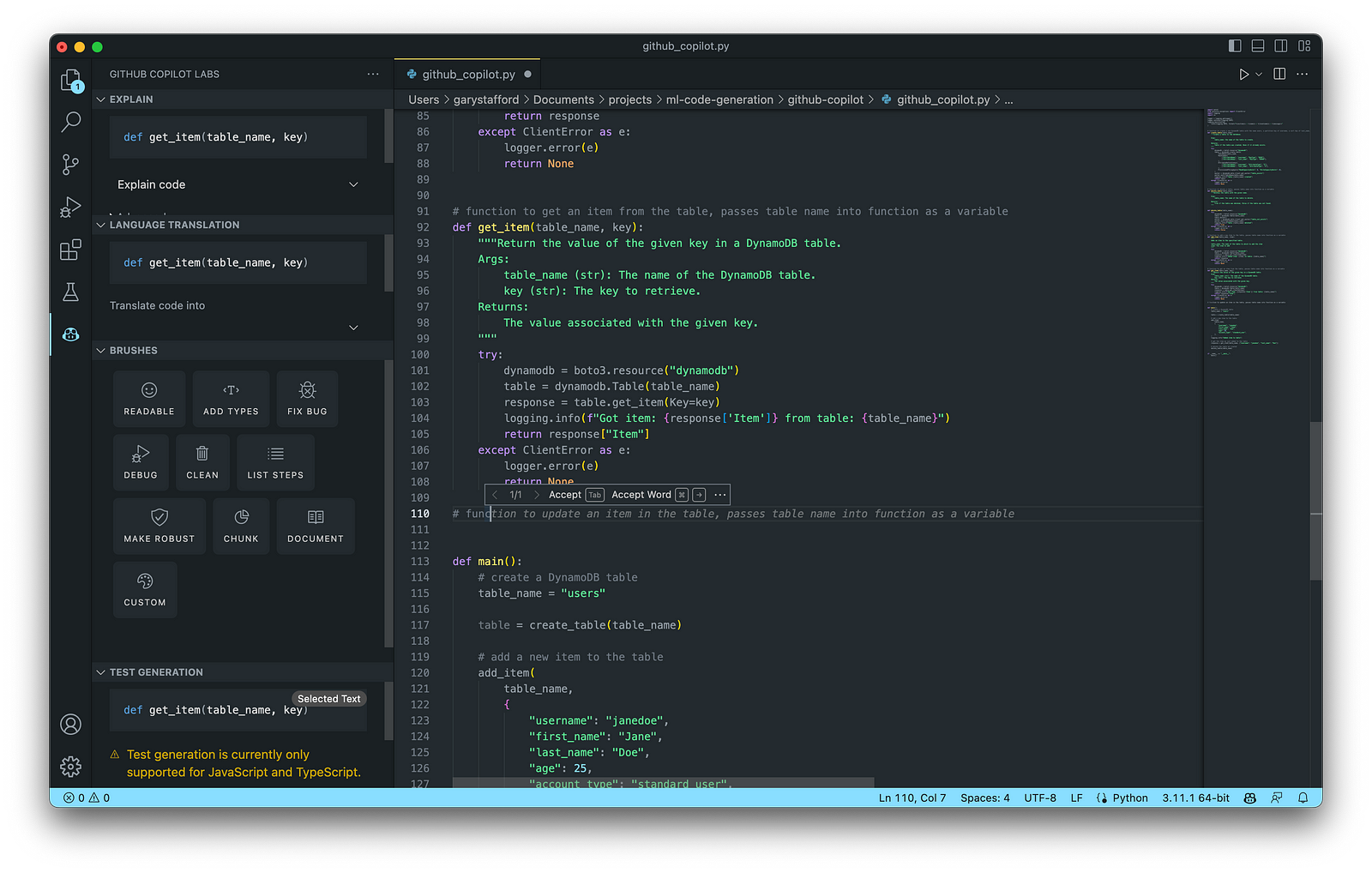
Copilot also has an extensive collapsible navigation bar, which can explain the code, offer language translations, provide access to Copilot’s Brushes, and can even generate tests. My only disappointment with Copilot is that test generation is currently only supported for JavaScript and TypeScript, given Python’s enormous popularity.

Below, we see an example of Copilot’s ability to generate whole-line code completion (line 14) and full-function code completion (lines 15–34) based on the existing code context and syntax (lines 1–14).

Copilot Results
Given my development knowledge and experience with similar tools, the final code results with Copilot were accurate, error-free, and well-formed. I found Copilot’s additional features, especially those found on its collapsible navigation bar, extended its functionality well beyond most similar generative AI tools I tested. Lastly, I felt Copilot’s ability to learn from the existing code context and syntax was superior to other tools.
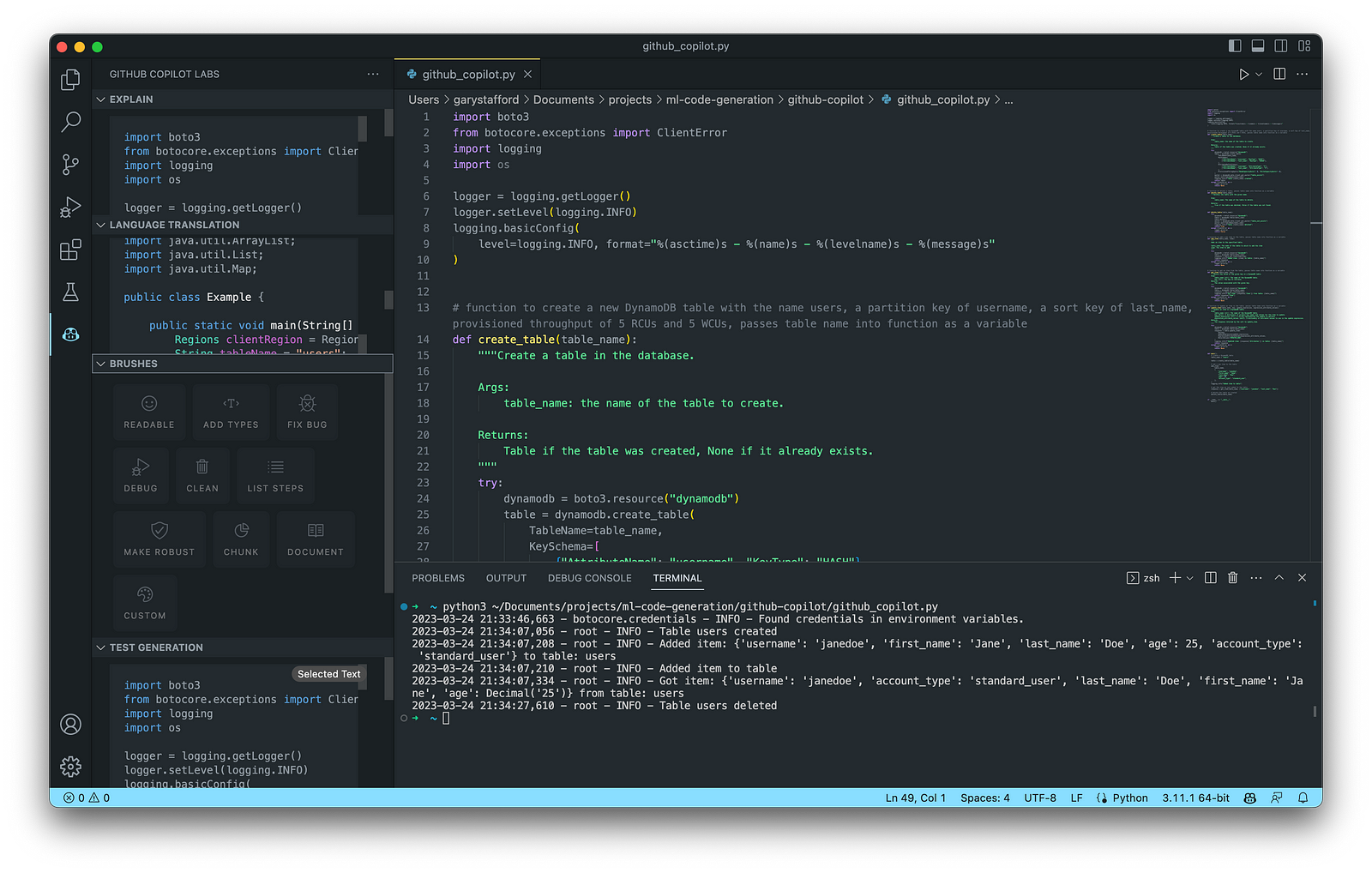
Amazon CodeWhisperer (Preview)

According to AWS, Amazon CodeWhisperer, announced in June 2022, “is a general purpose, machine learning-powered code generator that provides you with code recommendations, in real time. As you write code, CodeWhisperer automatically generates suggestions based on your existing code and comments. Your personalized recommendations can vary in size and scope, ranging from a single line comment to fully formed functions.” CodeWhisperer code generation is powered by ML models trained on various data sources, including Amazon and open-source code.
CodeWhisperer currently supports Java, JavaScript, Python, C#, and TypeScript. Although these are some of the most popular languages, CodeWhisperer’s language support is limited compared to Tabnine and Copilot.
Getting Started with CodeWhisperer
It is easy to get started with CodeWhisperer. Unfortunately, final pricing is unavailable since Amazon CodeWhisperer is still in Preview as of late March 2023.

CodeWhisperer enhances your IDE using AWS Toolkit for JetBrains, AWS Toolkit for Visual Studio Code, AWS Lambda console, and AWS Cloud9. With Lambda and AWS Cloud9, the setup simply involves activating CodeWhisperer within the IDE.

Getting started with CodeWhisperer requires installing the AWS Toolkit if applicable, choosing your authentication method, and setting up your Builder ID, IAM Identity Center (AWS SSO), or IAM.

AWS Builder ID
According to the documentation, “the AWS Builder ID is a new personal profile for everyone who builds on AWS. Your AWS Builder ID provides access to tools and builder services on AWS, including Amazon CodeCatalyst and Amazon CodeWhisperer. You can keep your AWS Builder ID as you move between jobs, schools, or other organizations. AWS Builder IDs are free. You only pay for the AWS resources you consume in your AWS accounts.”

Using CodeWhisperer
Nearly identical to Copilot and Tabnine, Amazon CodeWhisperer provides real-time code recommendations in your IDE. Unique features of CodeWhisperer include first-class support for AWS APIs, built-in security scans, a code reference tracker, and responsible AI/ML, which avoids bias by filtering out code recommendations that might be considered biased and unfair.
Below, we see an example of CodeWhisperer’s ability to generate whole-line code completion (line 14) and full-function code completion (lines 15–27) based on the existing code context and syntax (lines 1–14).

CodeWhisperer Results
Like my feedback on Tabnine and Copilot, the final code results with CodeWhisperer were accurate, error-free, and well-formed. In addition, I appreciated the added benefits of CodeWhisperer’s Security Scan and Reference Tracker to identify significant code from other projects.

Other Options
There are many other tools in the Generative AI coding category of comparable quality to the six featured in this post. New tools are being released on an almost daily basis.
Google Bard
On March 21, 2023, Google announced limited access to Bard, an early experiment that lets you collaborate with generative AI. They are beginning with the U.S. and the U.K. and will expand to more countries and languages over time. Bard is based on Google’s LaMDA (Language Model for Dialogue Applications), a transformer-based model.

Azure OpenAI Service with GPT-4
On March 21, 2023, Microsoft announced that GPT-4 was available in preview in Azure OpenAI Service. Unfortunately, Azure OpenAI Service requires registration and is currently only available to approved enterprise customers and partners. To get started, if you are qualified, you must trudge through the 25-question form.

Replit Ghostwriter
California-based VC-backed startup Replit has put the power of Replit’s AI into the world’s most popular online IDE. Replit “automates away the repetitive parts of coding, so you can stay focused on making your creative vision a reality.” Replit announced Ghostwriter Chat, which allows you to chat with a coding AI directly in your IDE. It now has a proactive debugger and awareness of your project’s code.
Conclusion
In this post, we examined six popular generative AI-powered coding tools, including chat-based OpenAI ChatGPT, Microsoft’s all-new Bing Chat, and ChatSonic, as well as IDE-based GitHub Copilot, Amazon CodeWhisperer (Preview), and Tabnine. Each tool assisted with developing an identical program to complete everyday database tasks on AWS. We then compared and contrasted the ease of use and the resulting code accuracy and quality of the tools. The generative AI space is moving at a breakneck pace. Tools continue to rapidly improve their AI models, add new features, and adjust pricing.
Recommended References
- Article: Top Artificial Intelligence (AI) Tools That Can Generate Code To Help Programmers (Marktechpost)
- Article: Top 29 ChatGPT alternatives that will blow your mind in 2023 (Free & Paid) (Writesonic)
- Research Paper: Deep Learning for Source Code Modeling and Generation: Models, Applications and Challenges (Triet H. M. Le, Hao Chen, M. Ali Babar)
- Article: NLP vs. NLU vs. NLG: the differences between three natural language processing concepts (IBM)
- Article: Top 17 Generative AI-based Programming Tools (For Developers) (Toward AI)
🔔 To keep up with future content, follow Gary Stafford on LinkedIn.
This blog represents my viewpoints and not those of my employer, Amazon Web Services (AWS). All product names, logos, and brands are the property of their respective owners.
Developing a Multi-Account AWS Environment Strategy
Posted by Gary A. Stafford in AWS, Cloud, DevOps, Enterprise Software Development, Technology Consulting on March 11, 2023
Explore twelve common patterns for developing an effective and efficient multi-account AWS environment strategy

Introduction
Every company is different: its organizational structure, the length of time it has existed, how fast it has grown, the industries it serves, its product and service diversity, public or private sector, and its geographic footprint. This uniqueness is reflected in how it organizes and manages its Cloud resources. Just as no two organizations are exactly alike, the structure of their AWS environments is rarely identical.
Some organizations successfully operate from a single AWS account, while others manage workloads spread across dozens or even hundreds of accounts. The volume and purpose of an organization’s AWS accounts are a result of multiple factors, including length of time spent on AWS, Cloud maturity, organizational structure and complexity, sectors, industries, and geographies served, product and service mix, compliance and regulatory requirements, and merger and acquisition activity.
“By design, all resources provisioned within an AWS account are logically isolated from resources provisioned in other AWS accounts, even within your own AWS Organizations.” (AWS)
Working with industry peers, the AWS community, and a wide variety of customers, one will observe common patterns for how organizations separate environments and workloads using AWS accounts. These patterns form an AWS multi-account strategy for operating securely and reliably in the Cloud at scale. The more planning an organization does in advance to develop a sound multi-account strategy, the less the burden that is required to manage changes as the organization grows over time.
The following post will explore twelve common patterns for effectively and efficiently organizing multiple AWS accounts. These patterns do not represent an either-or choice; they are designed to be purposefully combined to form a multi-account AWS environment strategy for your organization.
Patterns
- Pattern 1: Single “Uber” Account
- Pattern 2: Non-Prod/Prod Environments
- Pattern 3: Upper/Lower Environments
- Pattern 4: SDLC Environments
- Pattern 5: Major Workload Separation
- Pattern 6: Backup
- Pattern 7: Sandboxes
- Pattern 8: Centralized Management and Governance
- Pattern 9: Internal/External Environments
- Pattern 10: PCI DSS Workloads
- Pattern 11: Vendors and Contractors
- Pattern 12: Mergers and Acquisitions
Patterns 1–8 are progressively more mature multi-account strategies, while Patterns 9–12 represent special use cases for supplemental accounts.
Multi-Account Advantages
According to AWS’s whitepaper, Organizing Your AWS Environment Using Multiple Accounts, the benefits of using multiple AWS accounts include the following:
- Group workloads based on business purpose and ownership
- Apply distinct security controls by environment
- Constrain access to sensitive data (including compliance and regulation)
- Promote innovation and agility
- Limit the scope of impact from adverse events
- Support multiple IT operating models
- Manage costs (budgeting and cost attribution)
- Distribute AWS Service Quotas (fka limits) and API request rate limits
As we explore the patterns for organizing your AWS accounts, we will see how and to what degree each of these benefits is demonstrated by that particular pattern.
AWS Control Tower
Discussions about AWS multi-account environment strategies would not be complete without mentioning AWS Control Tower. According to the documentation, “AWS Control Tower offers a straightforward way to set up and govern an AWS multi-account environment, following prescriptive best practices.” AWS Control Tower includes Landing zone, described as “a well-architected, multi-account environment based on security and compliance best practices.”
AWS Control Tower is prescriptive in the Shared accounts it automatically creates within its AWS Organizations’ organizational units (OUs). Shared accounts created by AWS Control Tower include the Management, Log Archive, and Audit accounts. The previous standalone AWS service, AWS Landing Zone, maintained slightly different required accounts, including Shared Services, Log Archive, Security, and optional Network accounts. Although prescriptive, AWS Control Tower is also flexible and relatively unopinionated regarding the structure of Member accounts. Member accounts can be enrolled or unenrolled in AWS Control Tower.
You can decide whether or not to implement AWS Organizations or AWS Control Tower to set up and govern your AWS multi-account environment. Regardless, you will still need to determine how to reflect your organization’s unique structure and requirements in the purpose and quantity of the accounts you create within your AWS environment.
Common Multi-Account Patterns
While working with peers, community members, and a wide variety of customers, I regularly encounter the following twelve patterns for organizing AWS accounts. As noted earlier, these patterns do not represent an either-or choice; they are designed to be purposefully combined to form a multi-account AWS environment strategy for your organization.
Pattern 1: Single “Uber” Account
Organizations that effectively implement Pattern 1: Single “Uber” Account organize and separate environments and workloads at the sub-account level. They often use Amazon Virtual Private Cloud (Amazon VPC), an AWS account-level construct, to organize and separate environments and workloads. They may also use Subnets (VPC-level construct) or AWS Regions and Availability Zones to further organize and separate environments and workloads.
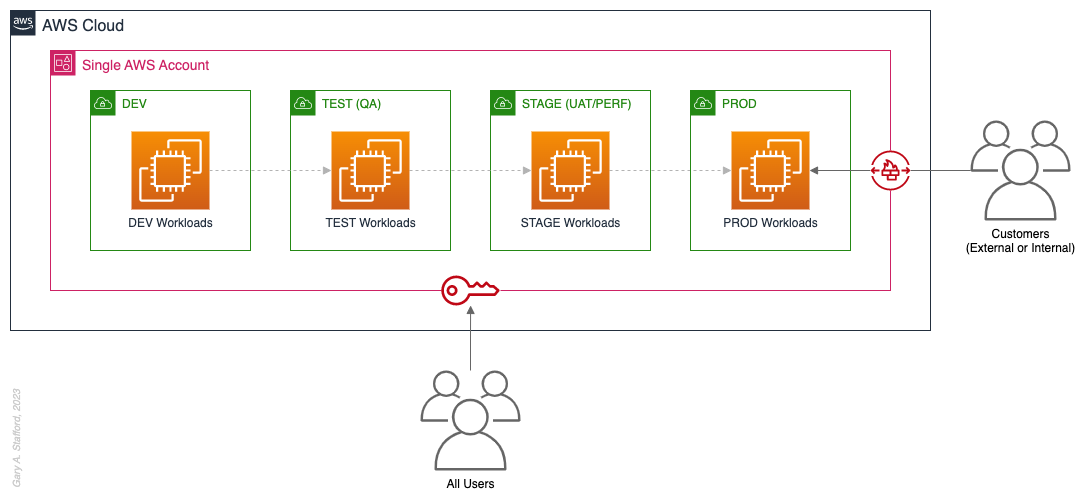
Pros
- Few, if any, significant advantages, especially for customer-facing workloads
Cons
- Decreased ability to limit the scope of impact from adverse events (widest blast radius)
- If the account is compromised, then all the organization’s workloads and data, possibly the entire organization, are potentially compromised (e.g., Ransomware attacks such as Encryptors, Lockers, and Doxware)
- Increased risk that networking or security misconfiguration could lead to unintended access to sensitive workloads and data
- Increased risk that networking, security, or resource management misconfiguration could lead to broad or unintended impairment of all workloads
- Decreased ability to perform team-, environment-, and workload-level budgeting and cost attribution
- Increased risk of resource depletion (soft and hard service quotas)
- Reduced ability to conduct audits and demonstrate compliance
Pattern 2: Non-Prod/Prod Environments
Organizations that effectively implement Pattern 2: Non-Prod/Prod Environments organize and separate non-Production workloads from Production (PROD) workloads using separate AWS accounts. Most often, they use Amazon VPCs within the non-Production account to separate workloads or Software Development Lifecycle (SDLC) environments, most often Development (DEV), Testing (TEST) or Quality Assurance (QA), and Staging (STAGE). Alternatively, these environments might also be designated as “N-1” (previous release), “N” (current release), “N+1” (next release), “N+2” (in development), and so forth, based on the currency of that version of that workload.
In some organizations, the Staging environment is used for User Acceptance Testing (UAT), performance (PERF) testing, and load testing before releasing workloads to Production. While in other organizations, STAGE, UAT, and PERF are each treated as separate environments at the account or VPC level.
Isolating Production workloads into their own account(s) and strictly limiting access to those workloads represents a significant first step in improving the overall maturity of your multi-account AWS environment strategy.
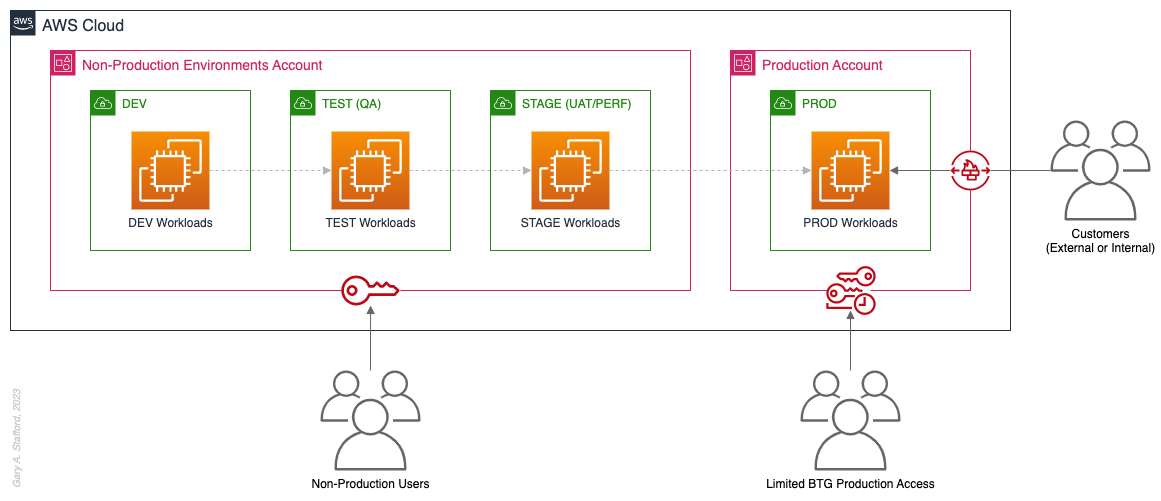
Pros
- Limits the scope of impact on Production as a result of adverse non-Production events (narrower blast radius)
- Logical separation and security of Production workloads and data
- Tightly control and limit access to Production, including the use of Break-the-Glass procedures (aka Break-glass or BTG); draws its name from “breaking the glass to pull a fire alarm”
- Eliminate the risk that non-Production networking or security misconfiguration could lead to unintended access to sensitive Production workloads and data
- Eliminate the chance that non-Production networking, security, or resource management misconfiguration could lead to broad or unintended impairment of Production workloads
- Conduct audits and demonstrate compliance with Production workloads
Cons
- If the Production account is compromised, then all the organization’s customer-facing workloads and data are potentially compromised
- Decreased ability to perform team-, environment-, and workload-level budgeting and cost attribution in the shared non-Production environment
- Increased risk of resource depletion (soft and hard service quotas) in the non-Production environment account
Pattern 3: Upper/Lower Environments
The next pattern, Pattern 3: Upper/Lower Environments, is a finer-grain variation of Pattern 2. With Pattern 3, we split all “Lower” environments into a single account and each “Upper” environment into its own account. In the software development process, initial environments, such as CI/CD for automated testing of code and infrastructure, Development, Test, UAT, and Performance, are called “Lower” environments. Conversely, later environments, such as Staging, Production, and even Disaster Recovery (DR), are called “Upper” environments. Upper environments typically require isolation for stability during testing or security for Production workloads and sensitive data.
Often, courser-grain patterns like Patterns 1–3 are carryovers from more traditional on-premises data centers, where compute, storage, network, and security resources were more constrained. Although these patterns can be successfully reproduced in the Cloud, they may not be optimal compared to more “cloud native” patterns, which provide improved separation of concerns.

Pros
- Limits the scope of impact on individual Upper environments as a result of adverse Lower environment events (narrower blast radius)
- Increased stability of Staging environment for critical UAT, performance, and load testing
- Logical separation and security of Production workloads and data
- Tightly control and limit access to Production, including the use of BTG
- Eliminate the risk that non-Production networking or security misconfiguration could lead to unintended access to sensitive Production workloads and data
- Eliminate the chance that non-Production networking, security, or resource management misconfiguration could lead to broad or unintended impairment of Production workloads
- Conduct audits and demonstrate compliance with Production workloads
Cons
- If the Production account is compromised, then all the organization’s customer-facing workloads and data are potentially compromised (e.g., Ransomware attack)
- Decreased ability to perform team-, environment-, and workload-level budgeting and cost attribution in the shared Lower environment
- Increased risk of resource depletion (soft and hard service quotas) in the Lower environment account
Pattern 4: SDLC Environments
The next pattern, Pattern 4: SDLC Environments, is a finer-grain variation of Pattern 3. With Pattern 4, we gain complete separation of each SDLC environment into its own AWS account. Using AWS services like AWS IAM Identity Center (fka AWS SSO), the Security team can enforce least-privilege permissions at an AWS Account level to individual groups of users, such as Developers, Testers, UAT, and Performance testers.
Based on my experience, Pattern 4 represents the minimal level of workload separation an organization should consider when developing its multi-account AWS environment strategy. Although Pattern 4 has a number of disadvantages, when combined with subsequent patterns and AWS best practices, this pattern begins to provide a scalable foundation for an organization’s growing workload portfolio.

Patterns, such as Pattern 4, not only apply to traditional software applications and services. These patterns can be applied to data analytics, AI/ML, IoT, media services, and similar workloads where separation of environments is required.

Pros
- Limits the scope of impact on one SDLC environment as a result of adverse events in another environment (narrower blast radius)
- Logical separation and security of Production workloads and data
- Increased stability of each SDLC environment
- Tightly control and limit access to Production, including the use of BTG
- Reduced risk that non-Production networking or security misconfiguration could lead to unintended access to sensitive Production workloads and data
- Reduced risk that non-Production networking, security, or resource management misconfiguration could lead to broad or unintended impairment of Production workloads
- Conduct audits and demonstrate compliance with Production workloads
- Increased ability to perform budgeting and cost attribution for each SDLC environment
- Reduced risk of resource depletion (soft and hard service limits) and IP conflicts and exhaustion within any single SDLC environment
Cons
- All workloads for each SDLC environment run within a single account, including Production, increasing the potential scope of impact from adverse events within that environment’s account
- If the Production account is compromised, then all the organization’s customer-facing workloads and data are potentially compromised
- Decreased ability to perform workload-level budgeting and cost attribution
Pattern 5: Major Workload Separation
The next pattern, Pattern 5: Major Workload Separation, is a finer-grain variation of Pattern 4. With Pattern 5, we separate each significant workload into its own separate SDLC environment account. The security team can enforce fine-grain least-privilege permissions at an AWS Account level to individual groups of users, such as Developers, Testers, UAT, and Performance testers, by their designated workload(s).
Pattern 5 has several advantages over the previous patterns. In addition to the increased workload-level security and reliability benefits, Pattern 5 can be particularly useful for organizations that operate significantly different technology stacks and specialized workloads, particularly at scale. Different technology stacks and specialized workloads often each have their own unique development, testing, deployment, and support processes. Isolating these types of workloads will help facilitate the support of multiple IT operating models.

Pros
- Limits the scope of impact on an individual workload as a result of adverse events from another workload or SDLC environment (narrowest blast radius)
- Increased ability to perform team-, environment-, and workload-level budgeting and cost attribution
Cons
- If the Production account is compromised, then all the organization’s customer-facing workloads and data are potentially compromised (e.g., Ransomware attack)
Pattern 6: Backup
In the earlier patterns, we mentioned that if the Production account were compromised, all the organization’s customer-facing workloads and data could be compromised. According to TechTarget, 2022 was a breakout year for Ransomware attacks. According to the US government’s CISA.gov website, “Ransomware is a form of malware designed to encrypt files on a device, rendering any files and the systems that rely on them unusable. Malicious actors then demand ransom in exchange for decryption.”
According to AWS best practices, one of the recommended preparatory actions to protect and recover from Ransomware attacks is backing up data to an alternate account using tools such as AWS Backup and an AWS Backup vault. Solutions such as AWS Backup protect and restore data regardless of how it was made inaccessible.
In Pattern 6: Backup, we create one or more Backup accounts to protect against unintended data loss or account compromise. In the example below, we have two Backup accounts, one for Production data and one for all non-Production data.

Pros
- If the Production account is compromised (e.g., Ransomware attacks such as Encryptors, Lockers, and Doxware), there are secure backups of data stored in a separate account, which can be used to restore or recreate the Production environment
Cons
- Few, if any, significant disadvantages when combined with previous patterns and AWS best-practices
Pattern 7: Sandboxes
The following pattern, Pattern 7: Sandboxes, supplements the previous patterns, designed to address the needs of an organization to allow individual users and teams to learn, build, experiment, and innovate on AWS without impacting the larger organization’s AWS environment. To quote the AWS blog, Best practices for creating and managing sandbox accounts in AWS, “Many organizations need another type of environment, one where users can build and innovate with AWS services that might not be permitted in production or development/test environments because controls have not yet been implemented.” Further, according to TechTarget, “a Sandbox is an isolated testing environment that enables users to run programs or open files without affecting the application, system, or platform on which they run.”
Due to the potential volume of individual user and team accounts, sometimes referred to as Sandbox accounts, mature infrastructure automation practices, cost controls, and self-service provisioning and de-provisioning of Sandbox accounts are critical capabilities for the organization.
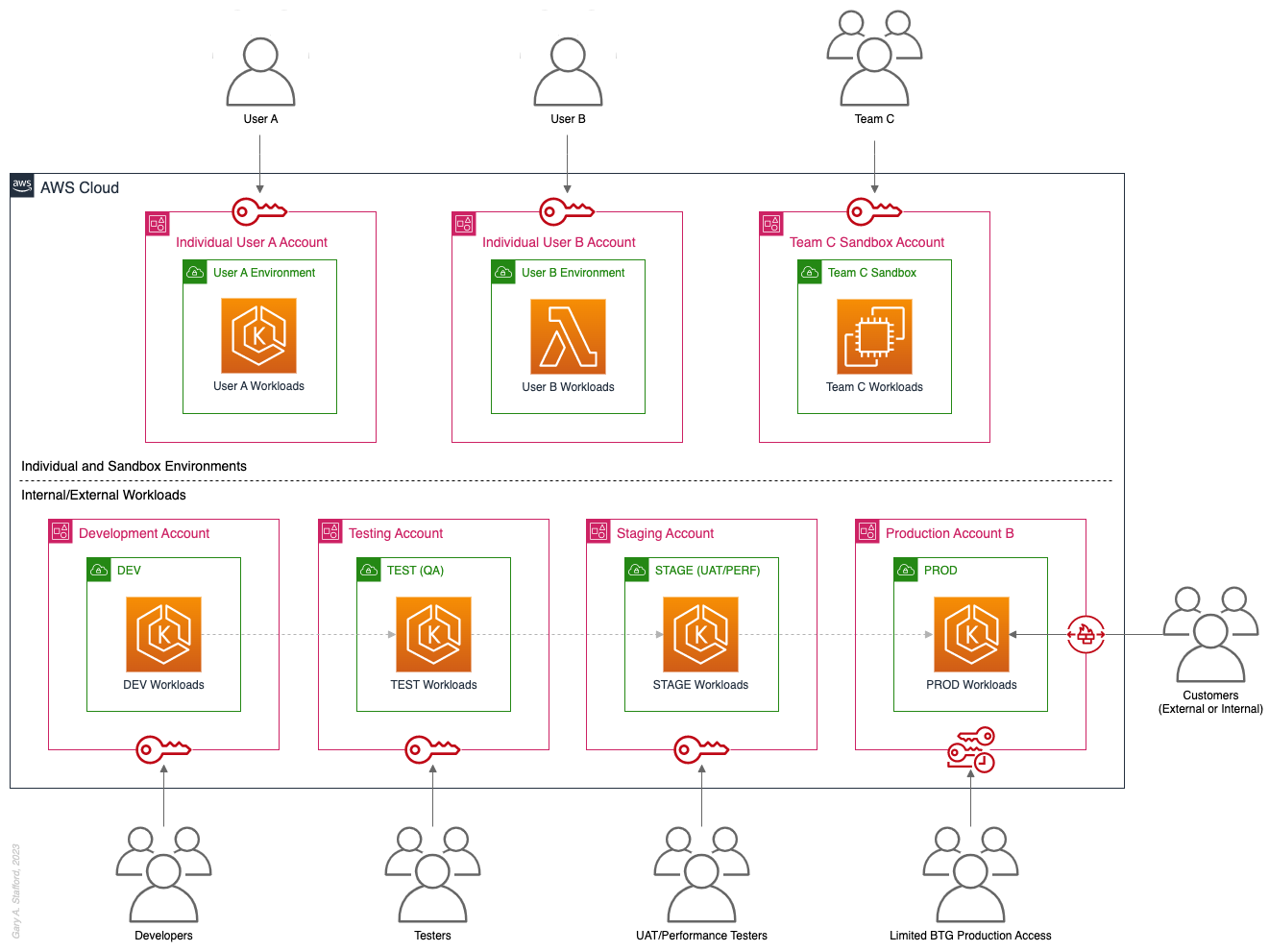
Pros
- Allow individual users and teams to learn, build, experiment, and innovate on AWS without impacting the rest of the organization’s AWS environment
Cons
- Without mature automation practices, cost controls, and self-service capabilities, managing multiple individual and team Sandbox accounts can become unwieldy and costly
Pattern 8: Centralized Management and Governance
We discussed AWS Control Tower at the beginning of this post. AWS Control Tower is prescriptive in creating Shared accounts within its AWS Organizations’ organizational units (OUs), including the Management, Log Archive, and Audit accounts. AWS encourages using AWS Control Tower to orchestrate multiple AWS accounts and services on your behalf while maintaining your organization’s security and compliance needs.
As exemplified in Pattern 8: Centralized Management and Governance, many organizations will implement centralized management whether or not they decide to implement AWS Control Tower. In addition to the Management account (payer account, fka master account), organizations often create centralized logging accounts, and centralized tooling (aka Shared services) accounts for functions such as CI/CD, IaC provisioning, and deployment. Another common centralized management account is a Security account. Organizations use this account to centralize the monitoring, analysis, notification, and automated mitigation of potential security issues within their AWS environment. The Security accounts will include services such as Amazon Detective, Amazon Inspector, Amazon GuardDuty, and AWS Security Hub.

Pros
- Increased ability to manage and maintain multiple AWS accounts with fewer resources
- Reduced duplication of management resources across accounts
- Increased ability to use automation and improve the consistency of processes and procedures across multiple accounts
Cons
- Few, if any, significant disadvantages when combined with previous patterns and AWS best-practices
Pattern 9: Internal/External Environments
The next pattern, Pattern 9: Internal/External Environments, focuses on organizations with internal operational systems (aka Enterprise systems) in the Cloud and customer-facing workloads. Pattern 9 separates internal operational systems, platforms, and workloads from external customer-facing workloads. For example, an organization’s divisions and departments, such as Sales and Marketing, Finance, Human Resources, and Manufacturing, are assigned their own AWS account(s). Pattern 9 allows the Security team to ensure that internal departmental or divisional users are isolated from users who are responsible for developing, testing, deploying, and managing customer-facing workloads.
Note that the diagram for Pattern 9 shows remote users who access AWS End User Computing (EUC) services or Virtual Desktop Infrastructure (VDI), such as Amazon WorkSpaces and Amazon AppStream 2.0. In this example, remote workers have secure access to EUC services provisioned in a separate AWS account, and indirectly, internal systems, platforms, and workloads.

Pros
- Separation of internal operational systems, platforms, and workloads from external customer-facing workloads
Cons
- Few, if any, significant disadvantages when combined with previous patterns and AWS best-practices
Pattern 10: PCI DSS Workloads
The next pattern, Pattern 10: PCI DSS Workloads, is a variation of previous patterns, which assumes the existence of Payment Card Industry Data Security Standard (PCI DSS) workloads and data. According to the AWS, “PCI DSS applies to entities that store, process, or transmit cardholder data (CHD) or sensitive authentication data (SAD), including merchants, processors, acquirers, issuers, and service providers. The PCI DSS is mandated by the card brands and administered by the Payment Card Industry Security Standards Council.”
According to AWS’s whitepaper, Architecting for PCI DSS Scoping and Segmentation on AWS, “By design, all resources provisioned within an AWS account are logically isolated from resources provisioned in other AWS accounts, even within your own AWS Organizations. Using an isolated account for PCI workloads is a core best practice when designing your PCI application to run on AWS.” With Pattern 10, we separate non-PCI DSS and PCI DSS Production workloads and data. The assumption is that only Production contains PCI DSS data. Data in lower environments is synthetically generated or sufficiently encrypted, masked, obfuscated, or tokenized.
Note that the diagram for Pattern 10 shows Administrators. Administrators with different spans of responsibility and access are present in every pattern, whether specifically shown or not.

Pros
- Increased ability to meet compliance requirements by separating non-PCI DSS and PCI DSS Production workloads
Cons
- Few, if any, significant disadvantages when combined with previous patterns and AWS best-practices
Pattern 11: Vendors and Contractors
The next pattern, Pattern 11: Vendors and Contractors, is focused on organizations that employ contractors or use third-party vendors who provide products and services that interact with their AWS-based environment. Like Pattern 10, Pattern 11 allows the Security team to ensure that contractor and vendor-based systems’ access to internal systems and customer-facing workloads is tightly controlled and auditable.
Vendor-based products and services are often deployed within an organization’s AWS environment without external means of ingress or egress. Alternatively, a vendor’s product or service may have a secure means of ingress from or egress to external endpoints. Such is the case with some SaaS products, which ship an organization’s data to an external aggregator for analytics or a security vendor’s product that pre-filters incoming data, external to the organization’s AWS environment. Using separate AWS accounts can improve an organization’s security posture and mitigate the risk of adverse events on the organization’s overall AWS environment.

Pros
- Ensure access to internal systems and customer-facing workloads by contractors and vendor-based systems is tightly controlled and auditable
Cons
- Few, if any, significant disadvantages when combined with previous patterns and AWS best-practices
Pattern 12: Mergers and Acquisitions
The next pattern, Pattern 12: Mergers and Acquisitions, is focused on managing the integration of external AWS accounts as a result of a merger or acquisition. This is a common occurrence, but the exact details of how best to handle the integration of two or more integrations depend on several factors. Factors include the required level of integration, for example, maintaining separate AWS Organizations, maintaining different AWS accounts, or merging resources from multiple accounts. Other factors that might impact account structure include changes in ownership or payer of acquired accounts, existing acquired cost-savings agreements (e.g., EDPs, PPAs, RIs, and Savings Plans), and AWS Marketplace vendor agreements. Even existing authentication and authorization methods of the acquiree versus the acquirer (e.g., AWS IAM Identity Center, Microsoft Active Directory (AD), Azure AD, and external identity providers (IdP) like Okta or Auth0).
The diagram for Pattern 12 attempts to show a few different M&A account scenarios, including maintaining separate AWS accounts for the acquirer and acquiree (e.g., acquiree’s Manufacturing Division account). If desired, the accounts can be kept independent but managed within the acquirer’s AWS Organizations’ organization. The diagram also exhibits merging resources from the acquiree’s accounts into the acquirer’s accounts (e.g., Sales and Marketing accounts). Resources will be migrated or decommissioned, and the account will be closed.

Pros
- Maintain separation between an acquirer and an acquiree’s AWS accounts within a single organization’s AWS environment
- Potentially consolidate and maximize cost-saving advantages of volume-related financial agreements and vendor licensing
Cons
- Migrating workloads between different organizations, depending on their complexity, requires careful planning and testing
- Consolidating multiple authentication and authorization methods requires careful planning and testing to avoid improper privileges
- Consolidating and optimizing separate licensing and cost-saving agreements between multiple organizations requires careful planning and an in-depth understanding of those agreements
Multi-Account AWS Environment Example
You can form an efficient and effective multi-account strategy for your organization by purposefully combining multiple patterns. Below is an example of combining the features of several patterns: Major Workload Separation, Backup, Sandboxes, Centralized Management and Governance, Internal/External Environments, and Vendors and Contractors.
According to AWS, “You can use AWS Organizations’ organizational units (OUs) to group accounts together to administer as a single unit. This greatly simplifies the management of your accounts.” If you decide to use AWS Organizations, each set of accounts associated with a pattern could correspond to an OU: Major Workload A, Major Workload B, Sandboxes, Backups, Centralized Management, Internal Environments, Vendors, and Contractors.

Conclusion
This post taught us twelve common patterns for effectively and efficiently organizing your AWS accounts. Instead of an either-or choice, these patterns are designed to be purposefully combined to form a multi-account strategy for your organization. Having a sound multi-account strategy will improve your security posture, maintain compliance, decrease the impact of adverse events on your AWS environment, and improve your organization’s ability to safely and confidently innovate and experiment on AWS.
Recommended References
- Establishing Your Cloud Foundation on AWS (AWS Whitepaper)
- Organizing Your AWS Environment Using Multiple Accounts (AWS Whitepaper)
- Building a Cloud Operating Model (AWS Whitepaper)
- AWS Security Reference Architecture (AWS Prescriptive Guidance)
- AWS Security Maturity Model (AWS Whitepaper)
- AWS re:Invent 2022 — Best practices for organizing and operating on AWS (YouTube Video)
- AWS Break Glass Role (GitHub)
🔔 To keep up with future content, follow Gary Stafford on LinkedIn.
This blog represents my viewpoints and not those of my employer, Amazon Web Services (AWS). All product names, logos, and brands are the property of their respective owners.
Brief Introduction to Observability on AWS
Posted by Gary A. Stafford in AWS, Cloud, DevOps on March 9, 2023
Explore the wide variety of Application Performance Monitoring (APM) and Observability options on AWS

APM and Observability
Observability is “the extent to which the internal states of a system can be inferred from externally available data” (Gartner). The three pillars of observability data are metrics, logs, and traces. Application Performance Monitoring (APM), a term commonly associated with observability, is “software that enables the observation and analysis of application health, performance, and user experience” (Gartner).
Additional features often associated with APM and observability products and services include the following (in alphabetical order):
- Advanced Threat Protection (ATP)
- Endpoint Detection and Response (EDR)
- Incident Detection Response (IDR)
- Infrastructure Performance Monitoring (IPM)
- Network Device Monitoring (NDM)
- Network Performance Monitoring (NPM)
- OpenTelemetry (OTel)
- Operational (or Operations) Intelligence Platform
- Predictive Monitoring (predictive analytics / predictive modeling)
- Real User Monitoring (RUM)
- Security Information and Event Management (SIEM)
- Security Orchestration, Automation, and Response (SOAR)
- Synthetic Monitoring (directed monitoring / synthetic testing)
- Threat Visibility and Risk Scoring
- Unified Security and Observability Platform
- User Behavior Analytics (UBA)
Not all features are offered by all vendors. Most vendors tend to specialize in one or more areas. Determining which features are essential to your organization before choosing a solution is vital.
AI/ML
Given the growing volume and real-time nature of observability telemetry, many vendors have started incorporating AI and ML into their products and services to improve correlation, anomaly detection, and mitigation capabilities. Understand how these features can reduce operational burden, enhance insights, and simplify complexity.
Decision Factors
APM and observability tooling choices often come down to a “Build vs. Buy” decision for organizations. In the Cloud, this usually means integrating several individual purpose-built products and services, self-managed open-source projects, or investing in an end-to-end APM or unified observability platform. Other decision factors include the need for solutions to support:
- Hybrid cloud environments (on-premises/Cloud)
- Multi-cloud environments (Public Cloud, SaaS, Supercloud)
- Specialized workloads (e.g., Mainframes, HPC, VMware, SAP, SAS)
- Compliant workloads (e.g., PCI DSS, PII, GDPR, FedRAMP)
- Edge Computing and IoT/IIoT
- AI, ML, and Data Analytics monitoring (AIOps, MLOps, DataOps)
- SaaS observability (SaaS providers who offer monitoring to their end-users as part of their service offering)
- Custom log formats and protocols
Finally, the 5 V’s of big data: Velocity, Volume, Value, Variety, and Veracity, also influence the choice of APM and observability tooling. The real-time nature of the observability data, the sheer volume of the data, the source and type of data, and the sensitivity of the data, will all guide tooling choices based on features and cost.
Organizations can choose fully-managed native AWS services, AWS Partner products and services, often SaaS, self-managed open-source observability tooling, or a combination of options. Many AWS and Partner products and services are commercial versions of popular open-source software (COSS).
AWS Options
- Amazon CloudWatch: Observe and monitor resources and applications you run on AWS in real time. CloudWatch features include Amazon CloudWatch Container, Lambda, Contributor, and Application Insights.
- Amazon Managed Service for Prometheus (AMP): Prometheus-compatible service that monitors and provides alerts on containerized applications and infrastructure at scale. Prometheus is “an open-source systems monitoring and alerting toolkit originally built at SoundCloud.”
- Amazon Managed Grafana (AMG): Commonly paired with AMP, AMG is a fully managed service for Grafana, an open-source analytics platform that “enables you to query, visualize, alert on, and explore your metrics, logs, and traces wherever they are stored.”
- Amazon OpenSearch Service: Based on open-source OpenSearch, this service provides interactive log analytics and real-time application monitoring and includes OpenSearch Dashboards (comparable to Kibana). Recently announced new security analytics features provide threat monitoring, detection, and alerting capabilities.
- AWS X-Ray: Trace user requests through your application, viewed using the X-Ray service map console or integrated with CloudWatch using the CloudWatch console’s X-Ray Traces Service map.
Data Collection, Processing, and Forwarding
- AWS Distro for OpenTelemetry (ADOT): Open-source APIs, libraries, and agents to collect distributed traces and metrics for application monitoring. ADOT is an open-source distribution of OpenTelemetry, a “high-quality, ubiquitous, and portable telemetry [solution] to enable effective observability.”
- AWS for Fluent Bit: Fluent Bit image with plugins for both CloudWatch Logs and Kinesis Data Firehose. Fluent Bit is an open-source, “super fast, lightweight, and highly scalable logging and metrics processor and forwarder.”
Security-focused Monitoring
- AWS CloudTrail: Helps enable operational and risk auditing, governance, and compliance in your AWS environment. CloudTrail records events, including actions taken in the AWS Management Console, AWS Command Line Interface (CLI), AWS SDKs, and APIs.
Partner Options
According to the 2022 Gartner® Magic Quadrant™ — APM & Observability Report, leading vendors commonly used by AWS customers include the following (in alphabetical order):
- AppDynamics (Cisco)
- Datadog
- Dynatrace
- Elastic
- Honeycomb
- Instanta (IBM)
- logz.io
- New Relic
- Splunk
- Sumo Logic
Open Source Options
There are countless open-source observability projects to choose from, including the following (in alphabetical order):
- Elastic: Elastic Stack: Elasticsearch, Kibana, Beats, and Logstash
- Fluentd: Data collector for unified logging layer
- Grafana: Platform for monitoring and observability
- Jaeger: End-to-end distributed tracing
- Kiali: Configure, visualize, validate, and troubleshoot Istio Service Mesh
- Loki: Like Prometheus, but for logs
- OpenSearch: Scalable, flexible, and extensible software suite for search, analytics, and observability applications
- OpenTelemetry (OTel): Collection of tools, APIs, and SDKs used to instrument, generate, collect, and export telemetry data
- Prometheus: Monitoring system and time series database
- Zabbix: Single pane of glass view of your whole IT infrastructure stack
🔔 To keep up with future content, follow Gary Stafford on LinkedIn.
This blog represents my viewpoints and not those of my employer, Amazon Web Services (AWS). All product names, logos, and brands are the property of their respective owners.
Building Data Lakes on AWS with Kafka Connect, Debezium, Apicurio Registry, and Apache Hudi
Posted by Gary A. Stafford in Analytics, AWS, Cloud on February 28, 2023
Learn how to build a near real-time transactional data lake on AWS using a combination of Open Source Software (OSS) and AWS Services
Introduction
In the following post, we will explore one possible architecture for building a near real-time transactional data lake on AWS. The data lake will be built using a combination of open source software (OSS) and fully-managed AWS services. Red Hat’s Debezium, Apache Kafka, and Kafka Connect will be used for change data capture (CDC). In addition, Apache Spark, Apache Hudi, and Hudi’s DeltaStreamer will be used to manage the data lake. To complete our architecture, we will use several fully-managed AWS services, including Amazon RDS, Amazon MKS, Amazon EKS, AWS Glue, and Amazon EMR.

Source Code
The source code, configuration files, and a list of commands shown in this post are open-sourced and available on GitHub.
Kafka
According to the Apache Kafka documentation, “Apache Kafka is an open-source distributed event streaming platform used by thousands of companies for high-performance data pipelines, streaming analytics, data integration, and mission-critical applications.” For this post, we will use Apache Kafka as the core of our change data capture (CDC) process. According to Wikipedia, “change data capture (CDC) is a set of software design patterns used to determine and track the data that has changed so that action can be taken using the changed data. CDC is an approach to data integration that is based on the identification, capture, and delivery of the changes made to enterprise data sources.” We will discuss CDC in greater detail later in the post.
There are several options for Apache Kafka on AWS. We will use AWS’s fully-managed Amazon Managed Streaming for Apache Kafka (Amazon MSK) service. Alternatively, you could choose industry-leading SaaS providers, such as Confluent, Aiven, Redpanda, or Instaclustr. Lastly, you could choose to self-manage Apache Kafka on Amazon Elastic Compute Cloud (Amazon EC2) or Amazon Elastic Kubernetes Service (Amazon EKS).
Kafka Connect
According to the Apache Kafka documentation, “Kafka Connect is a tool for scalably and reliably streaming data between Apache Kafka and other systems. It makes it simple to quickly define connectors that move large collections of data into and out of Kafka. Kafka Connect can ingest entire databases or collect metrics from all your application servers into Kafka topics, making the data available for stream processing with low latency.”
There are multiple options for Kafka Connect on AWS. You can use AWS’s fully-managed, serverless Amazon MSK Connect. Alternatively, you could choose a SaaS provider or self-manage Kafka Connect yourself on Amazon Elastic Compute Cloud (Amazon EC2) or Amazon Elastic Kubernetes Service (Amazon EKS). I am not a huge fan of Amazon MSK Connect during development. In my opinion, iterating on the configuration for a new source and sink connector, especially with transforms and external registry dependencies, can be painfully slow and time-consuming with MSK Connect. I find it much faster to develop and fine-tune my sink and source connectors using a self-managed version of Kafka Connect. For Production workloads, you can easily port the configuration from the native Kafka Connect connector to MSK Connect. I am using a self-managed version of Kafka Connect for this post, running in Amazon EKS.

Debezium
According to the Debezium documentation, “Debezium is an open source distributed platform for change data capture. Start it up, point it at your databases, and your apps can start responding to all of the inserts, updates, and deletes that other apps commit to your databases.” Regarding Kafka Connect, according to the Debezium documentation, “Debezium is built on top of Apache Kafka and provides a set of Kafka Connect compatible connectors. Each of the connectors works with a specific database management system (DBMS). Connectors record the history of data changes in the DBMS by detecting changes as they occur and streaming a record of each change event to a Kafka topic. Consuming applications can then read the resulting event records from the Kafka topic.”
Source Connectors
We will use Kafka Connect along with three Debezium connectors, MySQL, PostgreSQL, and SQL Server, to connect to three corresponding Amazon Relational Database Service (RDS) databases and perform CDC. Changes from the three databases will be represented as messages in separate Kafka topics. In Kafka Connect terminology, these are referred to as Source Connectors. According to Confluent.io, a leader in the Kafka community, “source connectors ingest entire databases and stream table updates to Kafka topics.”
Sink Connector
We will stream the data from the Kafka topics into Amazon S3 using a sink connector. Again, according to Confluent.io, “sink connectors deliver data from Kafka topics to secondary indexes, such as Elasticsearch, or batch systems such as Hadoop for offline analysis.” We will use Confluent’s Amazon S3 Sink Connector for Confluent Platform. We can use Confluent’s sink connector without depending on the entire Confluent platform.
There is also an option to use the Hudi Sink Connector for Kafka, which promises to greatly simplify the processes described in this post. However, the RFC for this Hudi feature appears to be stalled. Last updated in August 2021, the RFC is still in the initial “Under Discussion” phase. Therefore, I would not recommend the connectors use in Production until it is GA (General Availability) and gains broader community support.
Securing Database Credentials
Whether using Amazon MSK Connect or self-managed Kafka Connect, you should ensure your database, Kafka, and schema registry credentials, and other sensitive configuration values are secured. Both MSK Connect and self-managed Kafka Connect can integrate with configuration providers that implement the ConfigProvider class interface, such as AWS Secrets Manager, HashiCorp Vault, Microsoft Azure Key Vault, and Google Cloud Secrets Manager.
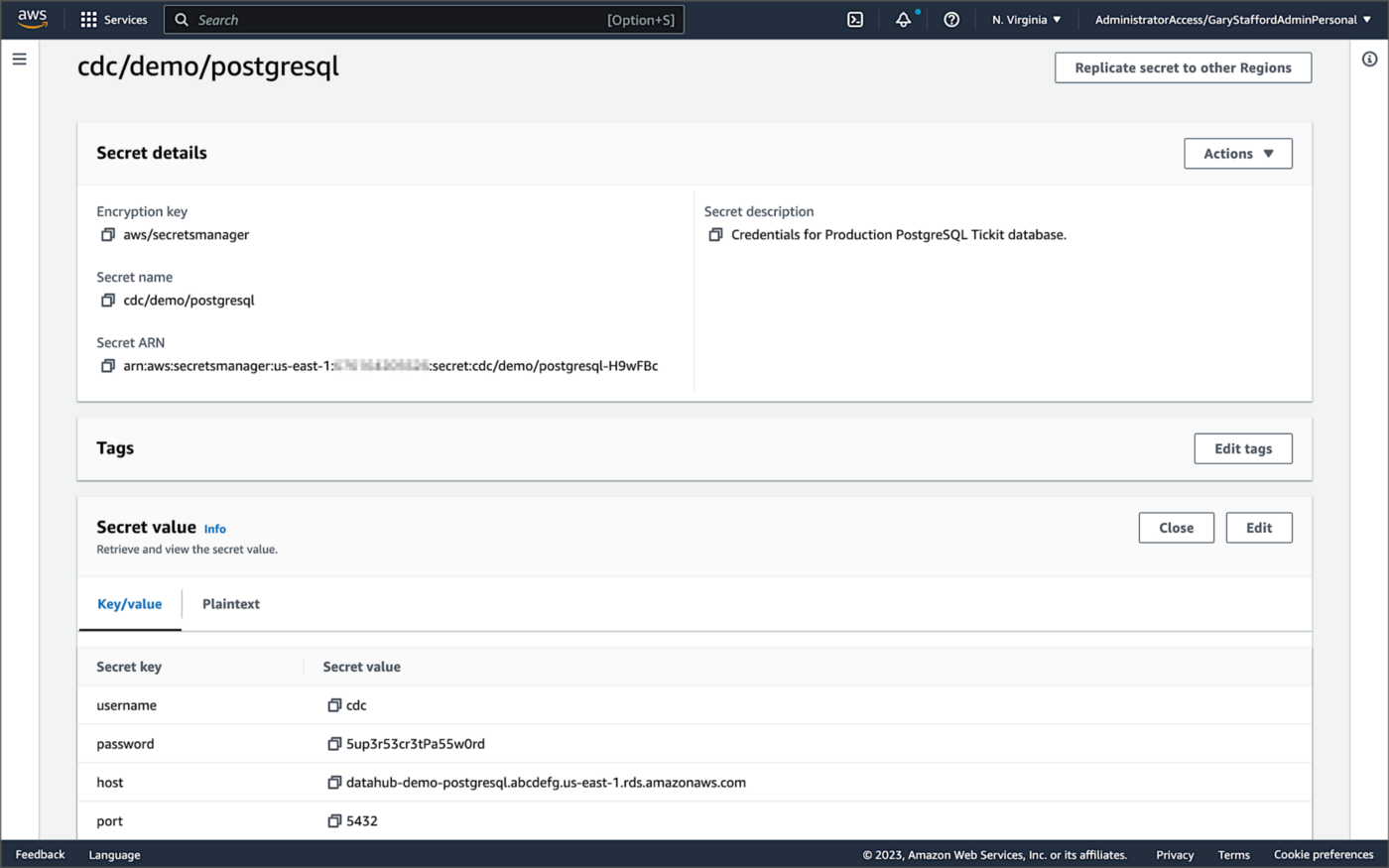
For self-managed Kafka Connect, I prefer Jeremy Custenborder’s kafka-config-provider-aws plugin. This plugin provides integration with AWS Secrets Manager. A complete list of Jeremy’s providers can be found on GitHub. Below is a snippet of the Secrets Manager configuration from the connect-distributed.properties files, which is read by Apache Kafka Connect at startup.
https://garystafford.medium.com/media/db6ff589c946fcd014546c56d0255fe5
Apache Avro
The message in the Kafka topic and corresponding objects in Amazon S3 will be stored in Apache Avro format by the CDC process. The Apache Avro documentation states, “Apache Avro is the leading serialization format for record data, and the first choice for streaming data pipelines.” Avro provides rich data structures and a compact, fast, binary data format.
Again, according to the Apache Avro documentation, “Avro relies on schemas. When Avro data is read, the schema used when writing it is always present. This permits each datum to be written with no per-value overheads, making serialization both fast and small. This also facilitates use with dynamic, scripting languages, since data, together with its schema, is fully self-describing.” When Avro data is stored in a file, its schema can be stored with it so any program may process files later.
Alternatively, the schema can be stored separately in a schema registry. According to Apicurio Registry, “in the messaging and event streaming world, data that are published to topics and queues often must be serialized or validated using a Schema (e.g. Apache Avro, JSON Schema, or Google protocol buffers). Schemas can be packaged in each application, but it is often a better architectural pattern to instead register them in an external system [schema registry] and then referenced from each application.”
Schema Registry
Several leading open-source and commercial schema registries exist, including Confluent Schema Registry, AWS Glue Schema Registry, and Red Hat’s open-source Apicurio Registry. In this post, we use a self-managed version of Apicurio Registry running on Amazon EKS. You can relatively easily substitute AWS Glue Schema Registry if you prefer a fully-managed AWS service.
Apicurio Registry
According to the documentation, Apicurio Registry supports adding, removing, and updating OpenAPI, AsyncAPI, GraphQL, Apache Avro, Google protocol buffers, JSON Schema, Kafka Connect schema, WSDL, and XML Schema (XSD) artifact types. Furthermore, content evolution can be controlled by enabling content rules, including validity and compatibility. Lastly, the registry can be configured to store data in various backend storage systems depending on the use case, including Kafka (e.g., Amazon MSK), PostgreSQL (e.g., Amazon RDS), and Infinispan (embedded).

Data Lake Table Formats
Three leading open-source, transactional data lake storage frameworks enable building data lake and data lakehouse architectures: Apache Iceberg, Linux Foundation Delta Lake, and Apache Hudi. They offer comparable features, such as ACID-compliant transactional guarantees, time travel, rollback, and schema evolution. Using any of these three data lake table formats will allow us to store and perform analytics on the latest version of the data in the data lake originating from the three Amazon RDS databases.
Apache Hudi
According to the Apache Hudi documentation, “Apache Hudi is a transactional data lake platform that brings database and data warehouse capabilities to the data lake.” The specifics of how the data is laid out as files in your data lake depends on the Hudi table type you choose, either Copy on Write (CoW) or Merge On Read (MoR).
Like Apache Iceberg and Delta Lake, Apache Hudi is partially supported by AWS’s Analytics services, including AWS Glue, Amazon EMR, Amazon Athena, Amazon Redshift Spectrum, and AWS Lake Formation. In general, CoW has broader support on AWS than MoR. It is important to understand the limitations of Apache Hudi with each AWS analytics service before choosing a table format.
If you are looking for a fully-managed Cloud data lake service built on Hudi, I recommend Onehouse. Born from the roots of Apache Hudi and founded by its original creator, Vinoth Chandar (PMC chair of the Apache Hudi project), the Onehouse product and its services leverage OSS Hudi to offer a data lake platform similar to what companies like Uber have built.
Hudi DeltaStreamer
Hudi also offers DeltaStreamer (aka HoodieDeltaStreamer) for streaming ingestion of CDC data. DeltaStreamer can be run once or continuously, using Apache Spark, similar to an Apache Spark Structured Streaming job, to capture changes to the raw CDC data and write that data to a different part of our data lake.
DeltaStreamer also works with Amazon S3 Event Notifications instead of running continuously. According to DeltaStreamer’s documentation, Amazon S3 object storage provides an event notification service to post notifications when certain events happen in your S3 bucket. AWS will put these events in Amazon Simple Queue Service (Amazon SQS). Apache Hudi provides an S3EventsSource that can read from Amazon SQS to trigger and process new or changed data as soon as it is available on Amazon S3.
Sample Data for the Data Lake
The data used in this post is from the TICKIT sample database. The TICKIT database represents the backend data store for a platform that brings buyers and sellers of tickets to entertainment events together. It was initially designed to demonstrate Amazon Redshift. The TICKIT database is a small database with approximately 425K rows of data across seven tables: Category, Event, Venue, User, Listing, Sale, and Date.
A data lake most often contains data from multiple sources, each with different storage formats, protocols, and connection methods. To simulate these data sources, I have separated the TICKIT database tables to represent three typical enterprise systems, including a commercial off-the-shelf (COTS) E-commerce platform, a custom Customer Relationship Management (CRM) platform, and a SaaS-based Event Management System (EMS). Each simulated system uses a different Amazon RDS database engine, including MySQL, PostgreSQL, and SQL Server.
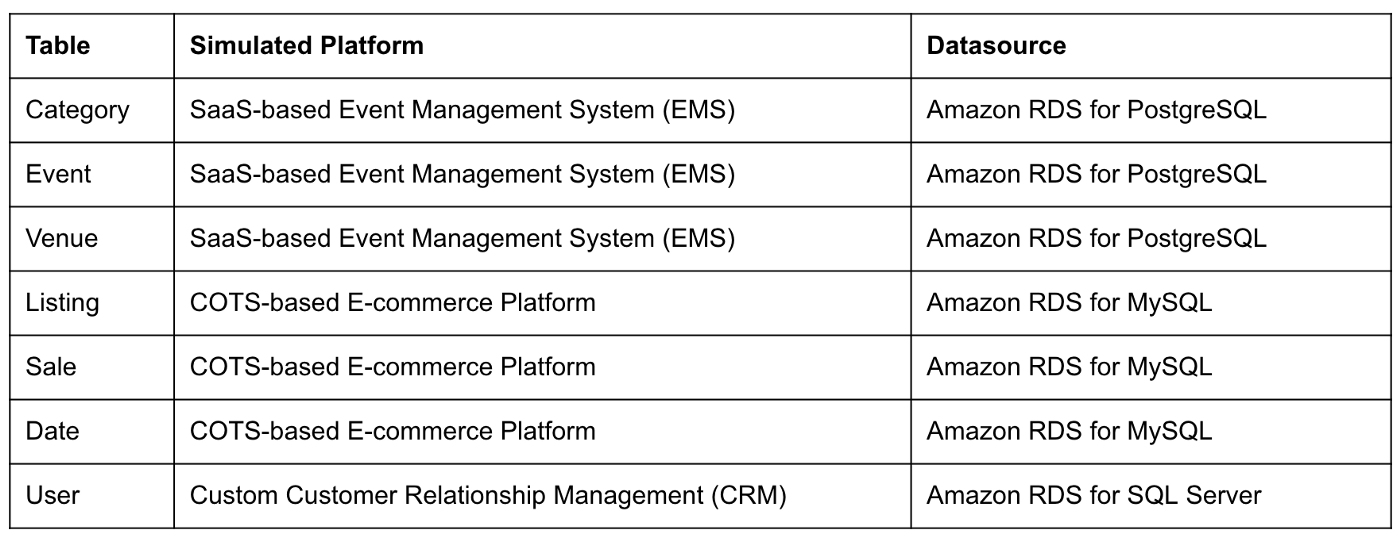
Enabling CDC for Amazon RDS
To use Debezium for CDC with Amazon RDS databases, minor changes to the default database configuration for each database engine are required.
CDC for PostgreSQL
Debezium has detailed instructions regarding configuring CDC for Amazon RDS for PostgreSQL. Database parameters specify how the database is configured. According to the Debezium documentation, for PostgreSQL, set the instance parameter rds.logical_replication to 1 and verify that the wal_level parameter is set to logical. It is automatically changed when the rds.logical_replication parameter is set to 1. This parameter is adjusted using an Amazon RDS custom parameter group. According to the AWS documentation, “with Amazon RDS, you manage database configuration by associating your DB instances and Multi-AZ DB clusters with parameter groups. Amazon RDS defines parameter groups with default settings. You can also define your own parameter groups with customized settings.”

CDC for MySQL
Similarly, Debezium has detailed instructions regarding configuring CDC for MySQL. Like PostgreSQL, MySQL requires a custom DB parameter group.


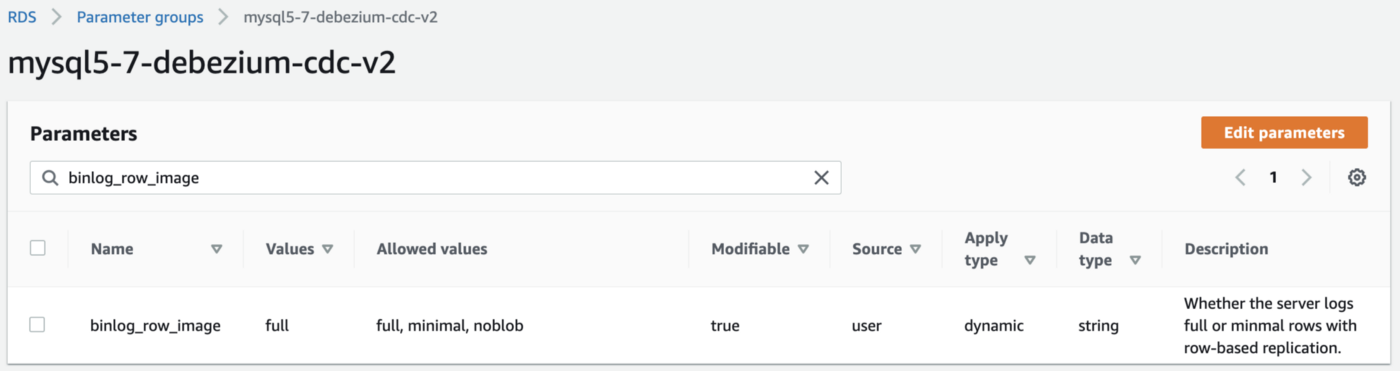
CDC for SQL Server
Lastly, Debezium has detailed instructions for configuring CDC with Microsoft SQL Server. Enabling CDC requires enabling CDC on the SQL Server database and table(s) and creating a new filegroup and associated file. Debezium recommends locating change tables in a different filegroup than you use for source tables. CDC is only supported with Microsoft SQL Server Standard Edition and higher; Express and Web Editions are not supported.
Kafka Connect Source Connectors
There is a Kafka Connect source connector for each of the three Amazon RDS databases, all of which use Debezium. CDC is performed, moving changes from the three databases into separate Kafka topics in Apache Avro format using the source connectors. The connector configuration is nearly identical whether you are using Amazon MSK Connect or a self-managed version of Kafka Connect.
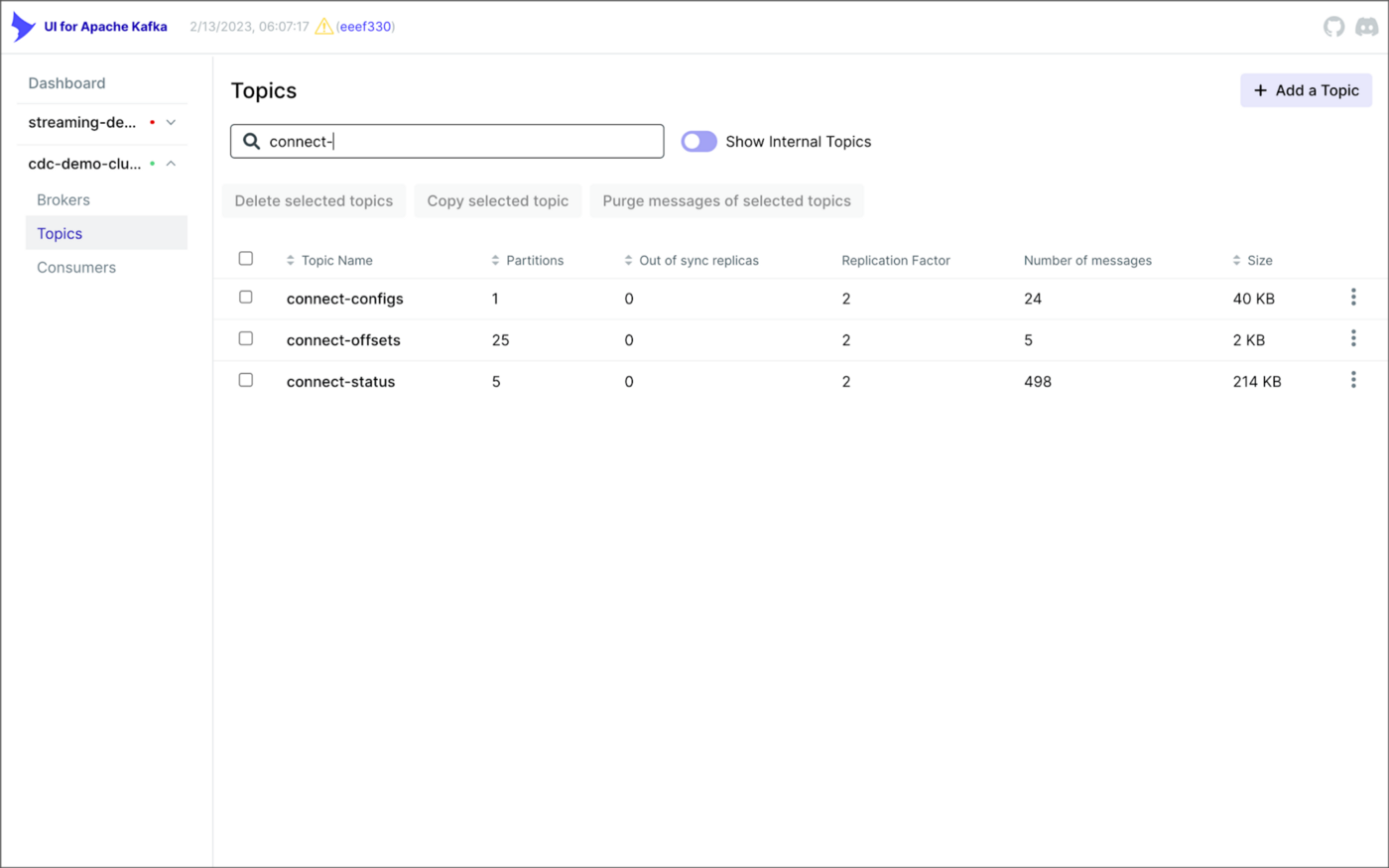
As shown above, I am using the UI for Apache Kafka by Provectus, self-managed on Amazon EKS, in this post.
PostgreSQL Source Connector
The source_connector_postgres_kafka_avro_tickit source connector captures all changes to the three tables in the Amazon RDS for PostgreSQL ticket database’s ems schema: category, event, and venue. These changes are written to three corresponding Kafka topics as Avro-format messages: tickit.ems.category, tickit.ems.event, and tickit.ems.venue. The messages are transformed by the connector using Debezium’s unwrap transform. Schemas for the messages are written to Apicurio Registry.
MySQL Source Connector
The source_connector_mysql_kafka_avro_tickit source connector captures all changes to the three tables in the Amazon RDS for PostgreSQL ecomm database: date, listing, and sale. These changes are written to three corresponding Kafka topics as Avro-format messages: tickit.ecomm.date, tickit.ecomm.listing, and tickit.ecomm.sale.
SQL Server Source Connector
Lastly, the source_connector_mssql_kafka_avro_tickit source connector captures all changes to the single user table in the Amazon RDS for SQL Server ticket database’s crm schema. These changes are written to a corresponding Kafka topic as Avro-format messages: tickit.crm.user.
If using a self-managed version of Kafka Connect, we can deploy, manage, and monitor the source and sink connectors using Kafka Connect’s RESTful API (Connect API).

Once the three source connectors are running, we should see seven new Kafka topics corresponding to the seven TICKIT database tables from the three Amazon RDS database sources. There will be approximately 425K messages consuming 147 MB of space in Avro’s binary format.
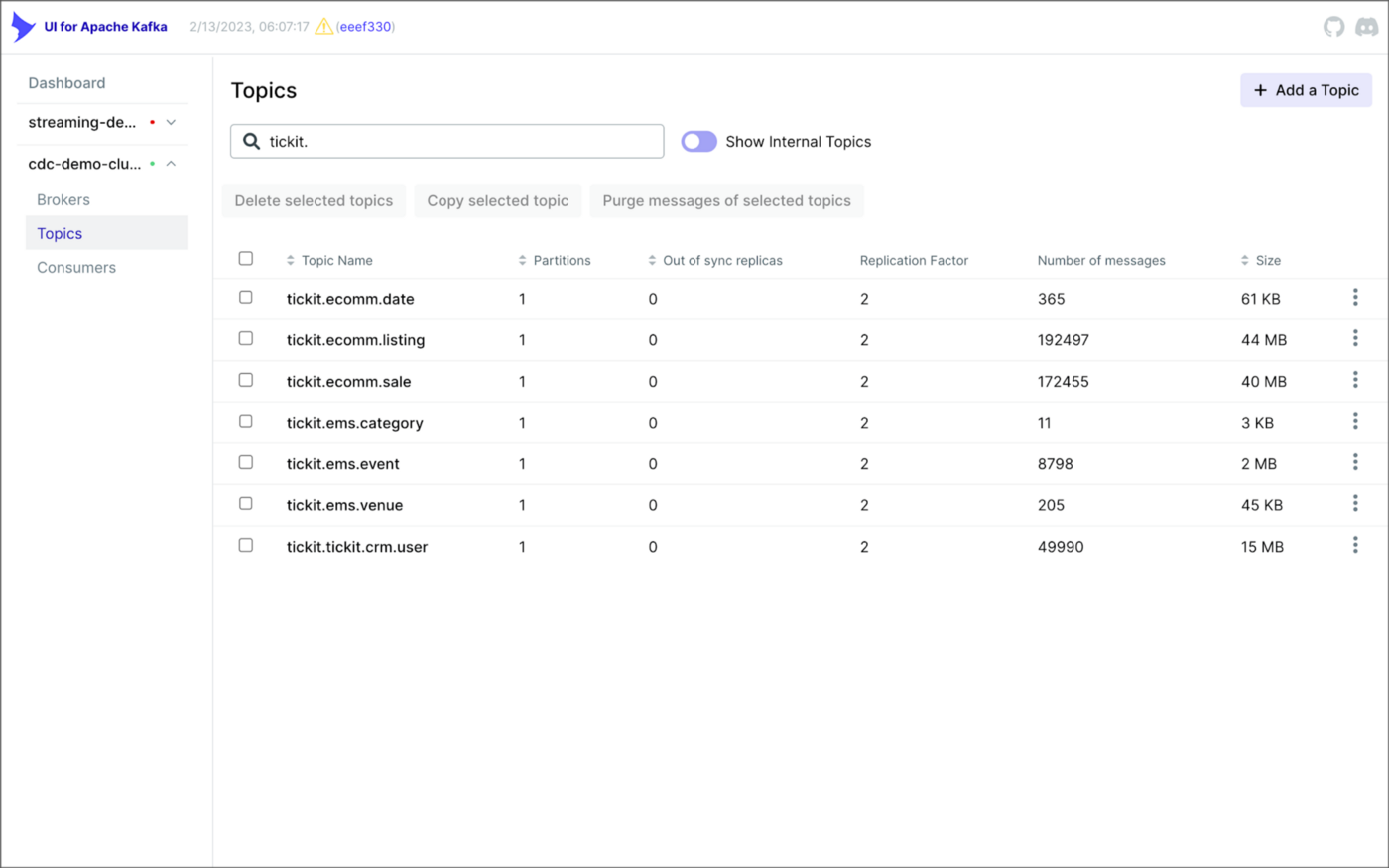
Avro’s binary format and the use of a separate schema ensure the messages consume minimal Kafka storage space. For example, the average message Value (payload) size in the tickit.ecomm.sale topic is a minuscule 98 Bytes. Resource management is critical when you are dealing with hundreds of millions or often billions of daily Kafka messages as a result of the CDC process.

From within the Apicurio Registry UI, we should now see Value schema artifacts for each of the seven Kafka topics.
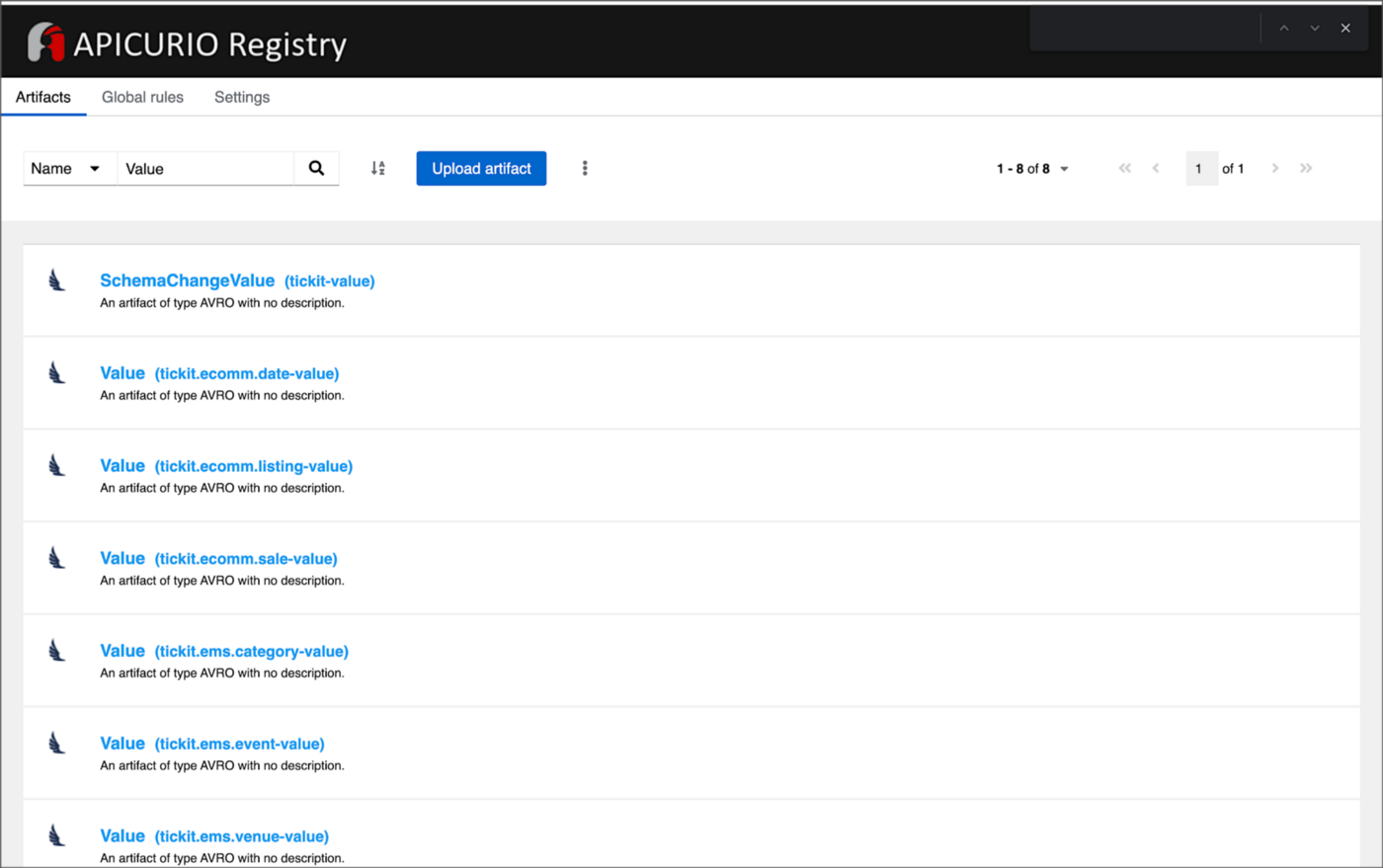
Also from within the Apicurio Registry UI, we can examine details of individual Value schema artifacts.

Kafka Connect Sink Connector
In addition to the three source connectors, there is a single Kafka Connect sink connector, sink_connector_kafka_s3_avro_tickit. The sink connector copies messages from the seven Kafka topics, all prefixed with tickit, to the Bronze area of our Amazon S3-based data lake.

The connector uses Confluent’s S3 Sink Connector (S3SinkConnector). Like Kafka, the connector writes all changes to Amazon S3 in Apache Avro format, with the schemas already stored in Apicurio Registry.
Once the sink connector and the three source connectors are running with Kafka Connect, we should see a series of seven subdirectories within the Bronze area of the data lake.

Confluent offers a choice of data partitioning strategies. The sink connector we have deployed uses the Daily Partitioner. According to the documentation, the io.confluent.connect.storage.partitioner.DailyPartitioner is equivalent to the TimeBasedPartitioner with path.format='year'=YYYY/'month'=MM/'day'=dd and partition.duration.ms=86400000 (one day for one S3 object in each daily directory). Below is an example of Avro files (S3 objects) containing changes to the sale table. The objects are partitioned by the year, month, and day they were written to the S3 bucket.

Message Transformation
Previously, while discussing the Kafka Connect source connectors, I mentioned that the connector transforms the messages using the unwrap transform. By default, the changes picked up by Debezium during the CDC process contain several additional Debezium-related fields of data. An example of an untransformed message generated by Debezium from the PostgreSQL sale table is shown below.
When the PostgreSQL connector first connects to a particular PostgreSQL database, it starts by performing a consistent snapshot of each database schema. All existing records are captured. Unlike an UPDATE ("op" : "u") or a DELETE ("op" : "d"), these initial snapshot messages, like the one shown below, represent a READ operation ("op" : "r"). As a result, there is no before data only after.
According to Debezium’s documentation, “Debezium provides a single message transformation [SMT] that crosses the bridge between the complex and simple formats, the UnwrapFromEnvelope SMT.” Below, we see the results of Debezium’s unwrap transform of the same message. The unwrap transform flattens the nested JSON structure of the original message, adds the __deleted field, and includes fields based on the list included in the source connector’s configuration. These fields are prefixed with a double underscore (e.g., __table).
In the next section, we examine Apache Hudi will manage the data lake. An example of the same message, managed with Hudi, is shown below. Note the five additional Hudi data fields, prefixed with _hoodie_.
Apache Hudi
With database changes flowing into the Bronze area of our data lake, we are ready to use Apache Hudi to provide data lake capabilities such as ACID transactional guarantees, time travel, rollback, and schema evolution. Using Hudi allows us to store and perform analytics on a specific time-bound version of the data in the data lake. Without Hudi, a query for a single row of data could return multiple results, such as an initial CREATE record, multiple UPDATE records, and a DELETE record.
DeltaStreamer
Using Hudi’s DeltaStreamer, we will continuously stream changes from the Bronze area of the data lake to a Silver area, which Hudi manages. We will run DeltaStreamer continuously, similar to an Apache Spark Structured Streaming job, using Apache Spark on Amazon EMR. Running DeltaStreamer requires using a spark-submit command that references a series of configuration files. There is a common base set of configuration items and a group of configuration items specific to each database table. Below, we see the base configuration, base.properties.
The base configuration is referenced by each of the table-specific configuration files. For example, below, we see the tickit.ecomm.sale.properties configuration file for DeltaStreamer.
To run DeltaStreamer, you can submit an EMR Step or use EMR’s master node to run a spark-submit command on the cluster. Below, we see an example of the DeltaStreamer spark-submit command using the Merge on Read (MoR) Hudi table type.
Next, we see the same example of the DeltaStreamer spark-submit command using the Copy on Write (CoW) Hudi table type.
For this post’s demonstration, we will run a single-table DeltaStreamer Spark job, with one process for each table using CoW. Alternately, Hudi offers HoodieMultiTableDeltaStreamer, a wrapper on top of HoodieDeltaStreamer, which enables the ingestion of multiple tables at a single time into Hudi datasets. Currently, HoodieMultiTableDeltaStreamer only supports sequential processing of tables to be ingested and COPY_ON_WRITE storage type.
Below, we see an example of three DeltaStreamer Spark jobs running continuously for the date, listing, and sale tables.

Once DeltaStreamer is up and running, we should see a series of subdirectories within the Silver area of the data lake, all managed by Apache Hudi.
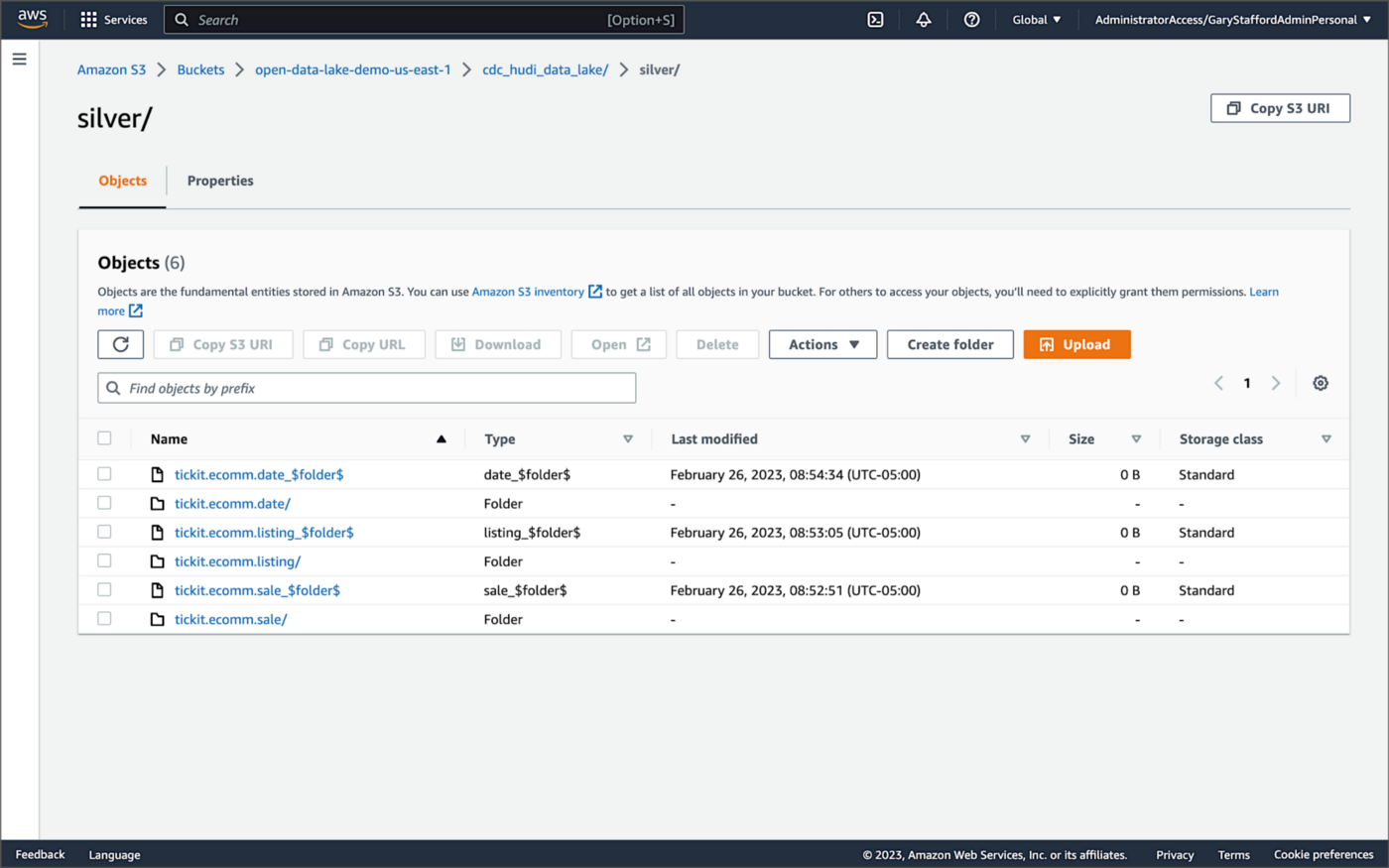
Within each subdirectory, partitioned by the table name, is a series of Apache Parquet files, along with other Apache Hudi-specific files. The specific folder structure and files depend on MoR or Cow.

AWS Glue Data Catalog
For this post, we are using an AWS Glue Data Catalog, an Apache Hive-compatible metastore, to persist technical metadata stored in the Silver area of the data lake, managed by Apache Hudi. The AWS Glue Data Catalog database, tickit_cdc_hudi, will be automatically created the first time DeltaStreamer runs.
Using DeltaStreamer with the table type of MERGE_ON_READ, there would be two tables in the AWS Glue Data Catalog database for each original table. According to Amazon’s EMR documentation, “Merge on Read (MoR) — Data is stored using a combination of columnar (Parquet) and row-based (Avro) formats. Updates are logged to row-based delta files and are compacted as needed to create new versions of the columnar files.” Hudi creates two tables in the Hive metastore for MoR, a table with the name that you specified, which is a read-optimized view (appended with _ro), and a table with the same name appended with _rt, which is a real-time view. You can query both tables.
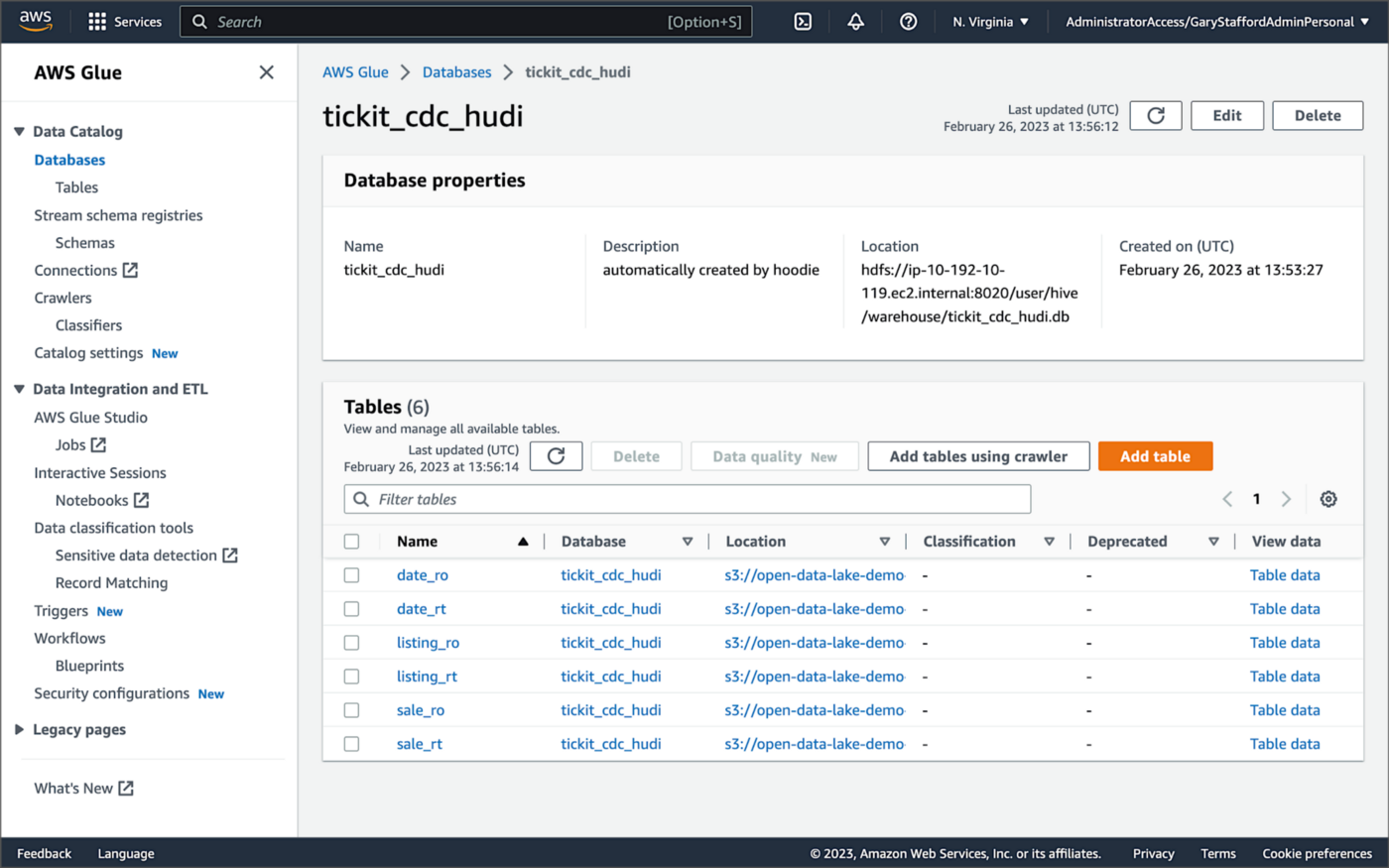
According to Amazon’s EMR documentation, “Copy on Write (CoW) — Data is stored in a columnar format (Parquet), and each update creates a new version of files during a write. CoW is the default storage type.” Using COPY_ON_WRITE with DeltaStreamer, there is only a single Hudi table in the AWS Glue Data Catalog database for each corresponding database table.
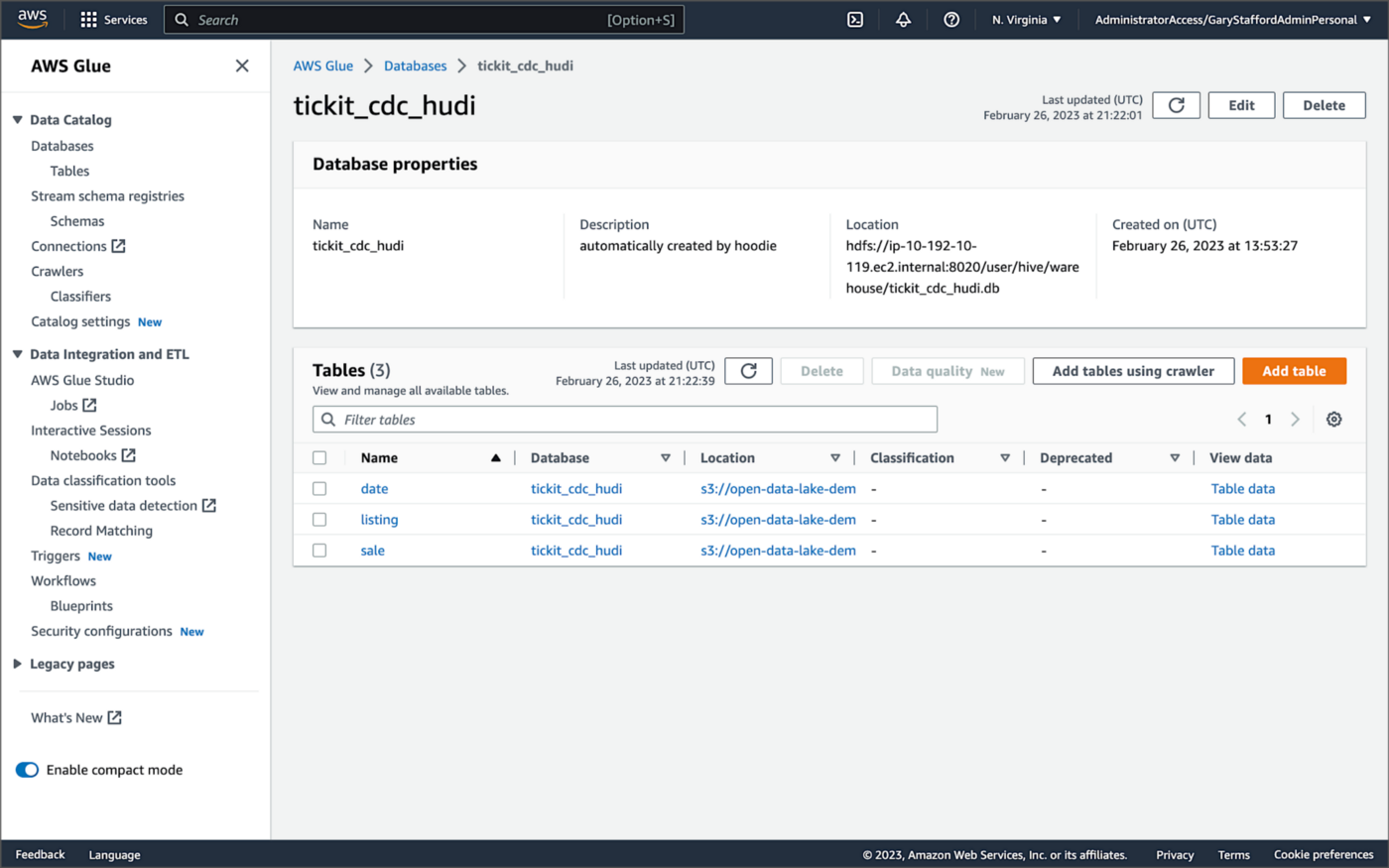
Examining an individual table in the AWS Glue Data Catalog database, we can see details like the table schema, location of underlying data in S3, input/output formats, Serde serialization library, and table partitions.
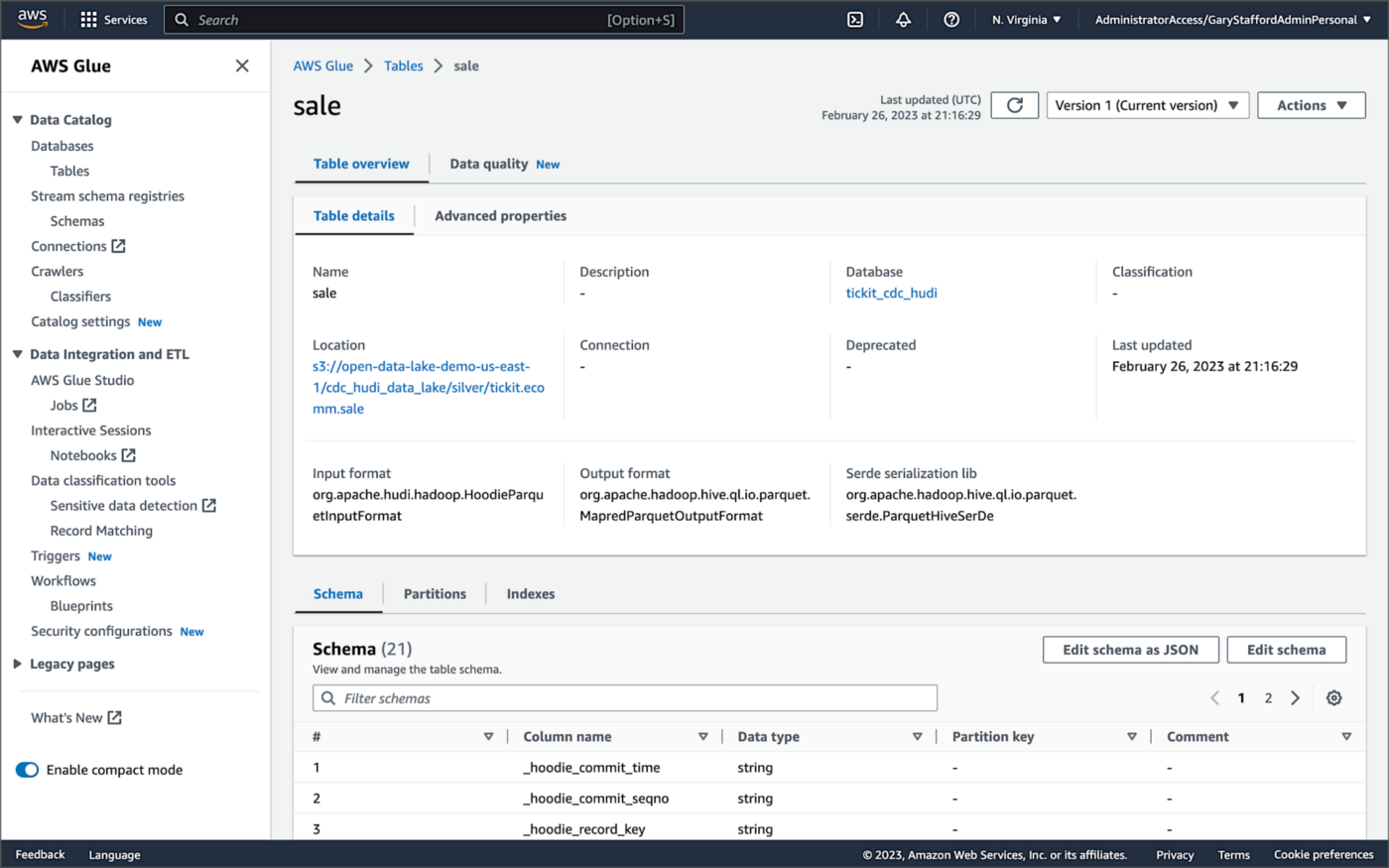
Database Changes and Data Lake
We are using Kafka Connect, Apache Hudi, and Hudi’s DeltaStreamer to maintain data parity between our databases and the data lake. Let’s look at an example of how a simple data change is propagated from a database to Kafka, then to the Bronze area of the data lake, and finally to the Hudi-managed Silver area of the data lake, written using the CoW table format.
First, we will make a simple update to a single record in the MySQL database’s sale table with the salesid = 200.
Almost immediately, we can see the change picked up by the Kafka Connect Debezium MySQL source connector.

Almost immediately, in the Bronze area of the data lake, we can see we have a new Avro-formatted file (S3 object) containing the updated record in the partitioned sale subdirectory.

If we examine the new object in the Bronze area of the data lake, we will see a row representing the updated record with the salesid = 200. Note how the operation is now an UPDATE versus a READ ("op" : "u").
Next, in the corresponding Silver area of the data lake, managed by Hudi, we should also see a new Parquet file that contains a record of the change. In this example, the file was written approximately 26 seconds after the original change to the database. This is the end-to-end time from database change to when the updated data is queryable in the data lake.
Similarly, if we examine the new object in the Silver area of the data lake, we will see a row representing the updated record with the salesid = 200. The record was committed approximately 15 seconds after the original database change.
Querying Hudi Tables with Amazon EMR
Using an EMR Notebook, we can query the updated database record stored in Amazon S3 using Hudi’s CoW table format. First, running a simple Spark SQL query for the record with salesid = 200, returns the latest record, reflecting the changes as indicated by the UPDATE operation value (u) and the _hoodie_commit_time = 2023-02-27 03:17:13.915 UTC.

Hudi Time Travel Query
We can also run a Hudi Time Travel Query, with an as.of.instance set to some arbitrary point in the future. Again, the latest record is returned, reflecting the changes as indicated by the UPDATE operation value (u) and the _hoodie_commit_time = 2023-02-27 03:17:13.915 UTC.

We can run the same Hudi Time Travel Query with an as.of.instance set to on or after the original records were written by DeltaStreamer (_hoodie_commit_time = 2023-02-27 03:13:50.642), but some arbitrary time before the updated record was written (_hoodie_commit_time = 2023-02-27 03:17:13.915). This time, the original record is returned as indicated by the READ operation value (r) and the _hoodie_commit_time = 2023-02-27 03:13:50.642. This is one of the strengths of Apache Hudi, the ability to query data in the present or at any point in the past.

Hudi Point in Time Query
We can also run a Hudi Point in Time Query, with a time range set from the beginning of time until some arbitrary point in the future. Again, the latest record is returned, reflecting the changes as indicated by the UPDATE operation value (u) and the _hoodie_commit_time = 2023-02-27 03:17:13.915.

We can run the same Hudi Point in Time Query but change the time range to two arbitrary time values, both before the updated record was written (_hoodie_commit_time = 2023-02-26 22:39:07.203). This time, the original record is returned as indicated by the READ operation value (r) and the _hoodie_commit_time of 2023-02-27 03:13:50.642. Again, this is one of the strengths of Apache Hudi, the ability to query data in the present or at any point in the past.

Querying Hudi Tables with Amazon Athena
We can also use Amazon Athena and run a SQL query against the AWS Glue Data Catalog database’s sale table to view the updated database record stored in Amazon S3 using Hudi’s CoW table format. The operation (op) column’s value now indicates an UPDATE (u).

How are Deletes Handled?
In this post’s architecture, deleted database records are signified with an operation ("op") field value of "d" for a DELETE and a __deleted field value of true.
https://itnext.io/media/f7c4ad62dcd45543fdf63834814a5aab
Back to our previous Jupyter notebook example, when rerunning the Spark SQL query, the latest record is returned, reflecting the changes as indicated by the DELETE operation value (d) and the _hoodie_commit_time = 2023-02-27 03:52:13.689.

Alternatively, we could use an additional Spark SQL filter statement to prevent deleted records from being returned (e.g., df.__op != "d").

Since we only did what is referred to as a soft delete in Hudi terminology, we can run a time travel query with an as.of.instance set to some arbitrary time before the deleted record was written (_hoodie_commit_time = 2023-02-27 03:52:13.689). This time, the original record is returned as indicated by the READ operation value (r) and the _hoodie_commit_time = 2023-02-27 03:13:50.642. We could also use a later as.of.instance to return the version of the record reflecting the the UPDATE operation value (u). This also applies to other query types such as point-in-time queries.

Conclusion
In this post, we learned how to build a near real-time transactional data lake on AWS using one possible architecture. The data lake was built using a combination of open source software (OSS) and fully-managed AWS services. Red Hat’s Debezium, Apache Kafka, and Kafka Connect were used for change data capture (CDC). In addition, Apache Spark, Apache Hudi, and Hudi’s DeltaStreamer were used to manage the data lake. To complete our architecture, we used several fully-managed AWS services, including Amazon RDS, Amazon MKS, Amazon EKS, AWS Glue, and Amazon EMR.
This blog represents my own viewpoints and not of my employer, Amazon Web Services (AWS). All product names, logos, and brands are the property of their respective owners.
Earning the AWS Certified Machine Learning — Specialty (MLS-C01) Certification
Posted by Gary A. Stafford in AWS, Cloud, Machine Learning on November 14, 2022
Introduction
Recently, I earned the AWS Certified Machine Learning — Specialty (MLS-C01) Certification, my ninth AWS certification. Since a few colleagues asked me about my preparation, I thought I would share it with the community, without divulging any details of the exam, of course.

Prerequisite Experience
Several AWS certifications can be earned with minimal to no hands-on AWS experience, but excellent short-term memorization skills. Although you will have technically earned the certification, you will certainly not be competent to practice the particular discipline. Certification does not equal qualification.
In my opinion, the AWS Certified Machine Learning — Specialty certification exam is not one of those where simple memorization of study materials, alone, will guarantee a passing score. If you lack practical experience in data science, machine learning, basic statistics, or data analytics on AWS, you will be challenged to pass this exam, no matter how much you cram.
Consider Data Analytics Certification First
To prepare for the Machine Learning — Specialty exam, I would strongly suggest first earning the AWS Certified Data Analytics — Specialty certification. According to the Machine Learning — Specialty exam’s content outline, “Domain 1: Data Engineering”, accounts for 20% of the exam’s score. Understanding the AWS Analytics services and how they integrate to form the most efficient data pipelines to feed your Machine Learning model training is a requirement for this portion of the exam’s questions. Preparing for the Data Analytics — Specialty certification will provide this adjacent domain knowledge:
- Amazon Athena
- Amazon EMR (pka Amazon Elastic MapReduce)
- Amazon IAM
- Amazon Kinesis Data Analytics, Data Firehose, Data Streams, Video Streams
- Amazon Redshift
- Amazon S3
- Amazon VPC
- AWS Data Pipeline
- AWS Glue Crawlers, Jobs, Data Catalog
- AWS Lambda
- AWS Step Functions

Study Materials
In my case, certification success was a result of practical experience, coursework, completing and reviewing the results of several practice exams, and taking lots of notes. The following is a list of the study materials I found most impactful:
Documentation
I reviewed the Amazon SageMaker and other AWS fully-managed AI/ML service documentation for my preparation.
Carefully review the Choose an Algorithm section of the Amazon SageMaker Developer Guide. According to the exam’s content outline, “Domain 3: Modeling” accounts for 36% of the exam’s score. Understand 1) recommended use cases for each of SageMaker’s built-in algorithms, 2) the algorithm’s required hyperparameters, and 3) the prescribed model evaluation metrics and tuning techniques. Built-in SageMaker algorithms most commonly covered in most training materials include:
- Tabular
- XGBoost (eXtreme Gradient Boosting)
- Linear Learner
- K-Nearest Neighbors (KNN)
- Factorization Machines
- Object2Vec
- Vision
- Image Classification
- Object Detection
- Semantic Segmentation
- Clustering
- K-Means
- Time-Series Forecast
- DeepAR
- Text Classification & Embedding
- BlazingText
- Text Transformation
- Sequence-to-Sequence (Seq2Seq)
- Text Topic Modeling
- Neural Topic Modeling (NTM)
- Latent Dirichlet Allocation (LDA)
- Dimensionality Reduction
- Principal Component Analysis (PCA)
- Anomaly Detection
- Random Cut Forest (RCF)
- IP Insights
AWS also uses Read the Docs. SageMaker’s Algorithm section is especially helpful with respect to preparing for the Machine Learning — Specialty exam: image processing, text processing, time-series processing, supervised learning, unsupervised learning, and feature engineering algorithms.
Along with algorithms, review SageMaker’s Deploy Models for Inference documentation. According to the exam’s content outline, “Domain 4: Machine Learning Implementation and Operations” accounts for 20% of the exam’s score. Understand SageMaker’s options for model serving, model versioning, deployment strategies, and endpoint monitoring.
Review the AWS fully managed AI/ML services Developer Guide documentation for the following services:
- Amazon Augmented AI
- Amazon CodeGuru
- Amazon Comprehend
- Amazon Forecast
- Amazon Fraud Detector
- Amazon Kendra
- Amazon Lex
- Amazon Personalize
- Amazon Polly
- Amazon Rekognition
- Amazon Textract
- Amazon Transcribe
- Amazon Translate
Understand the use cases for each of these services and most critically, how these managed services can be combined to create more complex AI/ML solutions. For example, building a near-real-time speech-to-speech translator with Amazon Transcribe, Amazon Translate, and Amazon Polly.
Online Courses
For my preparation, I completed three Udemy courses. Most of these online courses regularly go on sale and be purchased for $25 or less:
- AWS Certified Machine Learning Specialty 2022 — Hands On!, by Frank Kane and Stephane Maarek. Both Frank and Stephane are well-known across the industry and respected trainers. I recommend reviewing the algorithm, model evaluation, and high-level ML services sections more than once (Sections 5 and 6).
- AWS Certified Machine Learning Specialty (MLS-C01), by Chandra Lingam. Don’t get caught up in the nitty-gritty details of the Python code; focus on the higher-level machine learning principles. This course also contains a full-length practice exam.
- AWS Certified Machine Learning Specialty: 3 PRACTICE EXAMS, by Abhishek Singh. Nothing beats taking full-length practice exams and learning from your mistakes.
- Whizlabs’ AWS Certified Machine Learning Specialty Practice Tests. I completed a few of Whizlabs’ smaller practice exams, but, with limited time, I chose to complete Udemy’s full-length practice tests. Some of Whizlabs’ questions seemed off-topic to the exam outline and other training materials I reviewed.

Books
For my preparation, I read or re-read three books, two from Packt and one from O’Reilly:
- AWS Certified Machine Learning Specialty: MLS-C01 Certification Guide, by Somanath Nanda and Weslley Moura (Packt Publishing). I recommend this one if you only have time to read a single book.
- Practical Statistics for Data Scientists, 2nd Edition, by Peter Bruce, Andrew Bruce, Peter Gedeck (O’Reilly Media). According to the University of San Diego, “Statistics (or statistical analysis) is core to every machine learning algorithm.” This book covers many of the core statistical concepts behind Machine Learning, covered on the exam:
- BLEU
- Classification metrics: Precision-Recall Curve, ROC Curve, AUC
- Confusion Matrix: TP, FP, TN, FN, Accuracy, Precision, Recall (Sensitivity), Specificity, F1
- Correlated variables, Multicollinearity
- Distributions: Normal (Gaussian or “bell curve”), Bernoulli, Binomial, Poisson
- Elbow Method
- Ensemble Learning: Bagging, Boosting
- Euclidean Distance
- K-Fold Cross-Validation
- L1/L2 Regularization (lasso, alpha, ridge, lambda)
- Overfitting, Underfitting, High Bias, High Variance, Bias-Variance Tradeoff
- Plots: Histograms, Boxplots, Scatterplots
- Regression metrics: MAE, MSE, RMSE, R-squared, Adjusted R-squared
- Residuals
- SMOTE
- Standard Deviation, Three-Sigma/Empirical/68–95–99.7 Rule
- Z-score
- Python Machine Learning — Third Edition, by Sebastian Raschka and Vahid Mirjalili (Packt Publishing). Note that this book dives much deeper into the low-level statistical underpinnings of machine learning than is required for the exam, based on the exam outline. Again, don’t get caught up in the nitty-gritty details of Python; focus on the higher-level machine learning principles.
Scheduling the Exam
One last tip regarding when to take your exam. I have taken 15 AWS exams between nine AWS certifications and several recertifications. Although the Certified Machine Learning — Specialty exam is difficult, I found changing the time I sat the exam, greatly reduced my stress level. In the past, I took time off on a workday to complete exams, either in person or at home using online proctoring. I was preparing for the exam while frequently being interrupted by work-related items. For this exam, I chose to use online proctoring and took my exam at 6:00 AM on a Sunday morning. Up early, fresh, and full of energy, with no work- or family-related interruptions, no lawnmowers, dogs barking, or garbage trucks rumbling by, and no Internet bandwidth issues. I was done by 9:00 AM and eating breakfast with the family.
This blog represents my own viewpoints and not of my employer, Amazon Web Services (AWS). All product names, logos, and brands are the property of their respective owners.
Exploring Popular Open-source Stream Processing Technologies: Part 2 of 2
Posted by Gary A. Stafford in Analytics, Big Data, Java Development, Python, Software Development, SQL on September 26, 2022
A brief demonstration of Apache Spark Structured Streaming, Apache Kafka Streams, Apache Flink, and Apache Pinot with Apache Superset
Introduction
According to TechTarget, “Stream processing is a data management technique that involves ingesting a continuous data stream to quickly analyze, filter, transform or enhance the data in real-time. Once processed, the data is passed off to an application, data store, or another stream processing engine.” Confluent, a fully-managed Apache Kafka market leader, defines stream processing as “a software paradigm that ingests, processes, and manages continuous streams of data while they’re still in motion.”
This two-part post series and forthcoming video explore four popular open-source software (OSS) stream processing projects: Apache Spark Structured Streaming, Apache Kafka Streams, Apache Flink, and Apache Pinot.

This post uses the open-source projects, making it easier to follow along with the demonstration and keeping costs to a minimum. However, you could easily substitute the open-source projects for your preferred SaaS, CSP, or COSS service offerings.
Part Two
We will continue our exploration in part two of this two-part post, covering Apache Flink and Apache Pinot. In addition, we will incorporate Apache Superset into the demonstration to visualize the real-time results of our stream processing pipelines as a dashboard.
Demonstration #3: Apache Flink
In the third demonstration of four, we will examine Apache Flink. For this part of the post, we will also use the third of the three GitHub repository projects, flink-kafka-demo. The project contains a Flink application written in Java, which performs stream processing, incremental aggregation, and multi-stream joins.

New Streaming Stack
To get started, we need to replace the first streaming Docker Swarm stack, deployed in part one, with the second streaming Docker Swarm stack. The second stack contains Apache Kafka, Apache Zookeeper, Apache Flink, Apache Pinot, Apache Superset, UI for Apache Kafka, and Project Jupyter (JupyterLab).
https://programmaticponderings.wordpress.com/media/601efca17604c3a467a4200e93d7d3ff
The stack will take a few minutes to deploy fully. When complete, there should be ten containers running in the stack.
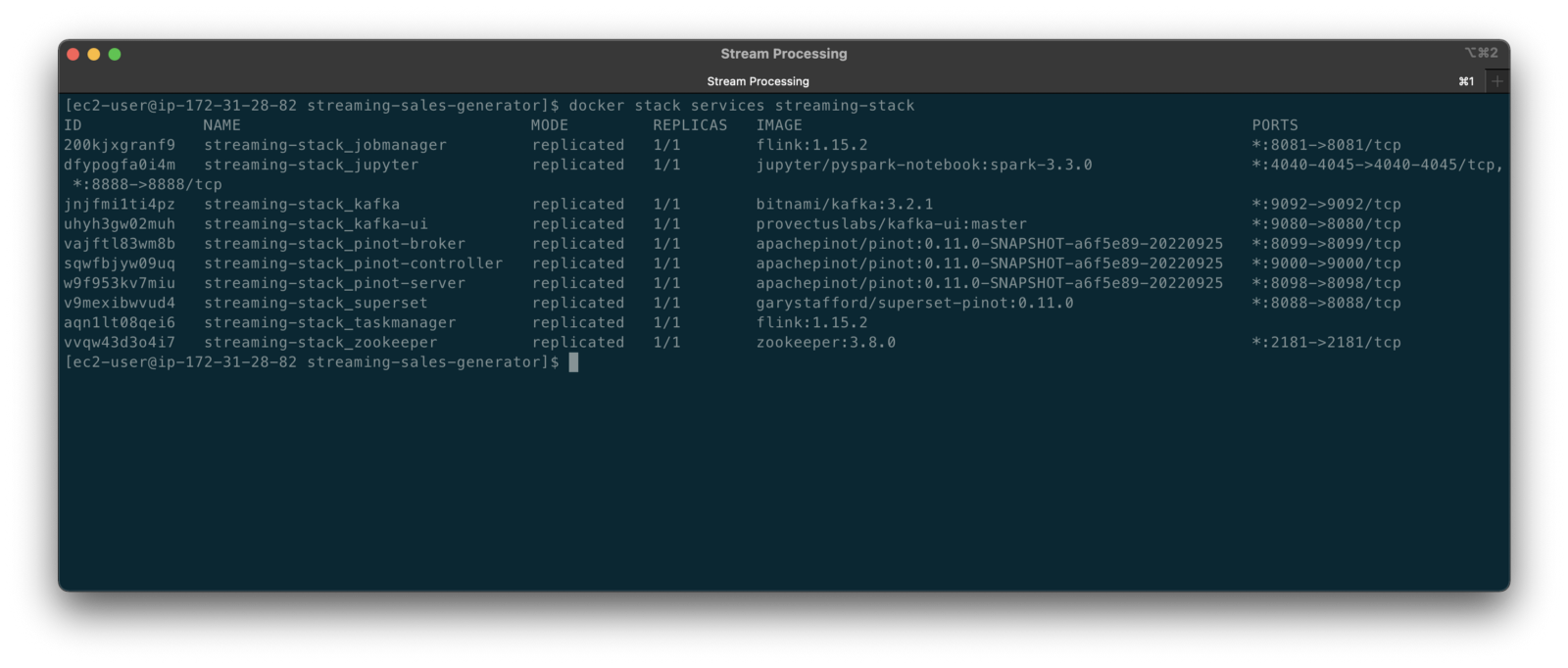
Flink Application
The Flink application has two entry classes. The first class, RunningTotals, performs an identical aggregation function as the previous KStreams demo.
The second class, JoinStreams, joins the stream of data from the demo.purchases topic and the demo.products topic, processing and combining them, in real-time, into an enriched transaction and publishing the results to a new topic, demo.purchases.enriched.
The resulting enriched purchases messages look similar to the following:
Running the Flink Job
To run the Flink application, we must first compile it into an uber JAR.
We can copy the JAR into the Flink container or upload it through the Apache Flink Dashboard, a browser-based UI. For this demonstration, we will upload it through the Apache Flink Dashboard, accessible on port 8081.
The project’s build.gradle file has preset the Main class (Flink’s Entry class) to org.example.JoinStreams. Optionally, to run the Running Totals demo, we could change the build.gradle file and recompile, or simply change Flink’s Entry class to org.example.RunningTotals.

Before running the Flink job, restart the sales generator in the background (nohup python3 ./producer.py &) to generate a new stream of data. Then start the Flink job.

To confirm the Flink application is running, we can check the contents of the new demo.purchases.enriched topic using the Kafka CLI.

Alternatively, you can use the UI for Apache Kafka, accessible on port 9080.

Demonstration #4: Apache Pinot
In the fourth and final demonstration, we will explore Apache Pinot. First, we will query the unbounded data streams from Apache Kafka, generated by both the sales generator and the Apache Flink application, using SQL. Then, we build a real-time dashboard in Apache Superset, with Apache Pinot as our datasource.

Creating Tables
According to the Apache Pinot documentation, “a table is a logical abstraction that represents a collection of related data. It is composed of columns and rows (known as documents in Pinot).” There are three types of Pinot tables: Offline, Realtime, and Hybrid. For this demonstration, we will create three Realtime tables. Realtime tables ingest data from streams — in our case, Kafka — and build segments from the consumed data. Further, according to the documentation, “each table in Pinot is associated with a Schema. A schema defines what fields are present in the table along with the data types. The schema is stored in Zookeeper, along with the table configuration.”
Below, we see the schema and config for one of the three Realtime tables, purchasesEnriched. Note how the columns are divided into three categories: Dimension, Metric, and DateTime.
To begin, copy the three Pinot Realtime table schemas and configurations from the streaming-sales-generator GitHub project into the Apache Pinot Controller container. Next, use a docker exec command to call the Pinot Command Line Interface’s (CLI) AddTable command to create the three tables: products, purchases, and purchasesEnriched.
To confirm the three tables were created correctly, use the Apache Pinot Data Explorer accessible on port 9000. Use the Tables tab in the Cluster Manager.

We can further inspect and edit the table’s config and schema from the Tables tab in the Cluster Manager.

The three tables are configured to read the unbounded stream of data from the corresponding Kafka topics: demo.products, demo.purchases, and demo.purchases.enriched.
Querying with Pinot
We can use Pinot’s Query Console to query the Realtime tables using SQL. According to the documentation, “Pinot provides a SQL interface for querying. It uses the [Apache] Calcite SQL parser to parse queries and uses MYSQL_ANSI dialect.”

With the generator still running, re-query the purchases table in the Query Console (select count(*) from purchases). You should notice the document count increasing each time you re-run the query since new messages are published to the demo.purchases topic by the sales generator.
If you do not observe the count increasing, ensure the sales generator and Flink enrichment job are running.

Table Joins?
It might seem logical to want to replicate the same multi-stream join we performed with Apache Flink in part three of the demonstration on the demo.products and demo.purchases topics. Further, we might presume to join the products and purchases realtime tables by writing a SQL statement in Pinot’s Query Console. However, according to the documentation, at the time of this post, version 0.11.0 of Pinot did not [currently] support joins or nested subqueries.
This current join limitation is why we created the Realtime table, purchasesEnriched, allowing us to query Flink’s real-time results in the demo.purchases.enriched topic. We will use both Flink and Pinot as part of our stream processing pipeline, taking advantage of each tool’s individual strengths and capabilities.
Note, according to the documentation for the latest release of Pinot on the main branch, “the latest Pinot multi-stage supports inner join, left-outer, semi-join, and nested queries out of the box. It is optimized for in-memory process and latency.” For more information on joins as part of Pinot’s new multi-stage query execution engine, read the documentation, Multi-Stage Query Engine.

demo.purchases.enriched topic in real-timeAggregations
We can perform real-time aggregations using Pinot’s rich SQL query interface. For example, like previously with Spark and Flink, we can calculate running totals for the number of items sold and the total sales for each product in real time.

We can do the same with the purchasesEnriched table, which will use the continuous stream of enriched transaction data from our Apache Flink application. With the purchasesEnriched table, we can add the product name and product category for richer results. Each time we run the query, we get real-time results based on the running sales generator and Flink enrichment job.

Query Options and Indexing
Note the reference to the Star-Tree index at the start of the SQL query shown above. Pinot provides several query options, including useStarTree (true by default).
Multiple indexing techniques are available in Pinot, including Forward Index, Inverted Index, Star-tree Index, Bloom Filter, and Range Index, among others. Each has advantages in different query scenarios. According to the documentation, by default, Pinot creates a dictionary-encoded forward index for each column.
SQL Examples
Here are a few examples of SQL queries you can try in Pinot’s Query Console:
Troubleshooting Pinot
If have issues with creating the tables or querying the real-time data, you can start by reviewing the Apache Pinot logs:
Real-time Dashboards with Apache Superset
To display the real-time stream of data produced results of our Apache Flink stream processing job and made queriable by Apache Pinot, we can use Apache Superset. Superset positions itself as “a modern data exploration and visualization platform.” Superset allows users “to explore and visualize their data, from simple line charts to highly detailed geospatial charts.”
According to the documentation, “Superset requires a Python DB-API database driver and a SQLAlchemy dialect to be installed for each datastore you want to connect to.” In the case of Apache Pinot, we can use pinotdb as the Python DB-API and SQLAlchemy dialect for Pinot. Since the existing Superset Docker container does not have pinotdb installed, I have built and published a Docker Image with the driver and deployed it as part of the second streaming stack of containers.
First, we much configure the Superset container instance. These instructions are documented as part of the Superset Docker Image repository.
Once the configuration is complete, we can log into the Superset web-browser-based UI accessible on port 8088.
Pinot Database Connection and Dataset
Next, to connect to Pinot from Superset, we need to create a Database Connection and a Dataset.
The SQLAlchemy URI is shown below. Input the URI, test your connection (‘Test Connection’), make sure it succeeds, then hit ‘Connect’.
Next, create a Dataset that references the purchasesEnriched Pinot table.

purchasesEnriched Pinot tableModify the dataset’s transaction_time column. Check the is_temporal and Default datetime options. Lastly, define the DateTime format as epoch_ms.
transaction_time columnBuilding a Real-time Dashboard
Using the new dataset, which connects Superset to the purchasesEnriched Pinot table, we can construct individual charts to be placed on a dashboard. Build a few charts to include on your dashboard.
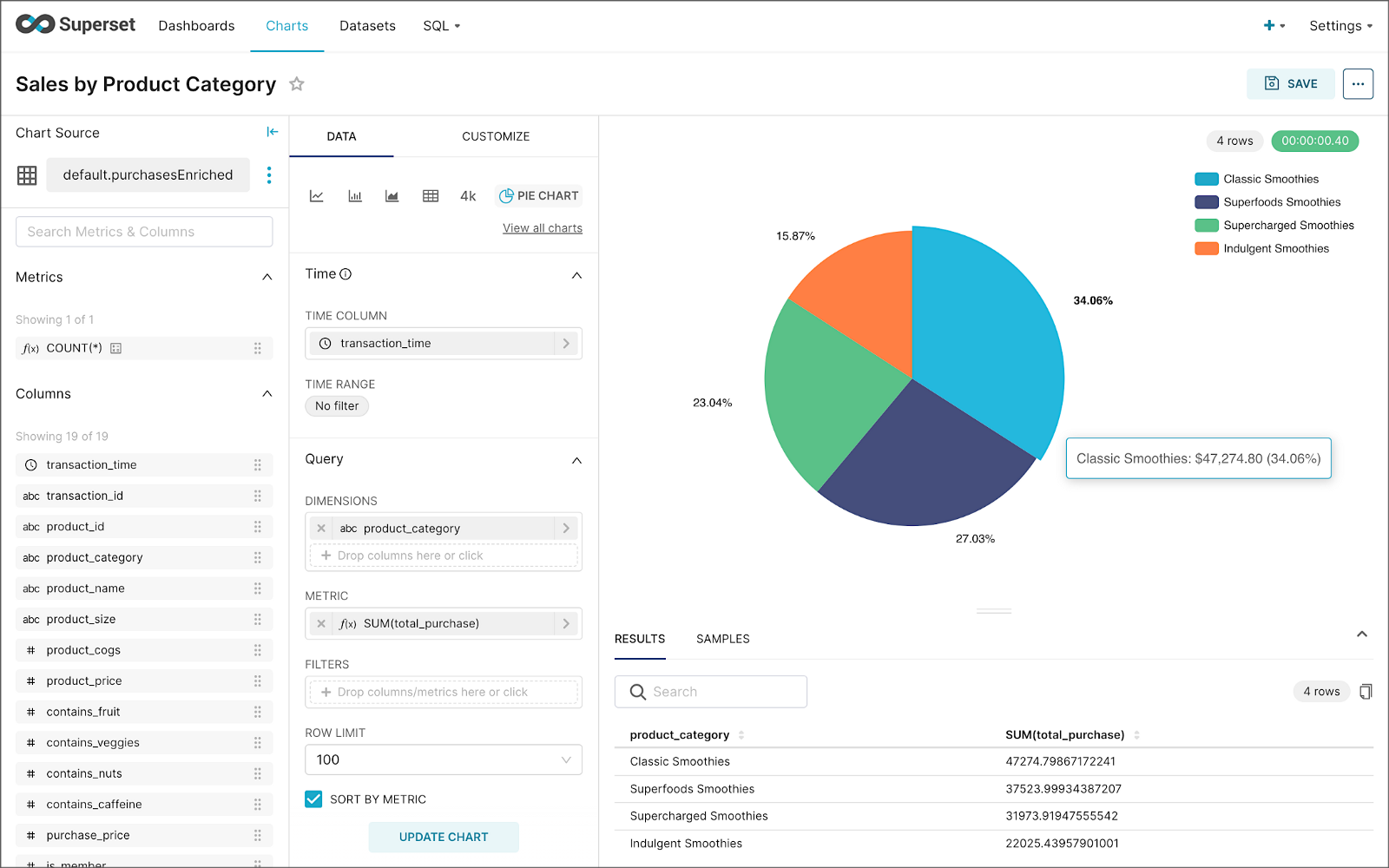
Create a new Superset dashboard and add the charts and other elements, such as headlines, dividers, and tabs.

We can apply a refresh interval to the dashboard to continuously query Pinot and visualize the results in near real-time.
Conclusion
In this two-part post series, we were introduced to stream processing. We explored four popular open-source stream processing projects: Apache Spark Structured Streaming, Apache Kafka Streams, Apache Flink, and Apache Pinot. Next, we learned how we could solve similar stream processing and streaming analytics challenges using different streaming technologies. Lastly, we saw how these technologies, such as Kafka, Flink, Pinot, and Superset, could be integrated to create effective stream processing pipelines.
This blog represents my viewpoints and not of my employer, Amazon Web Services (AWS). All product names, logos, and brands are the property of their respective owners. All diagrams and illustrations are the property of the author unless otherwise noted.